

The generalization error (test error) of a ML system can be always decomposed into two parts:
underfitting. Solution: increase the complexity/expressiveness of your ML algorithm!overfitting. Solution: decrease the model complexity or add strong regularization.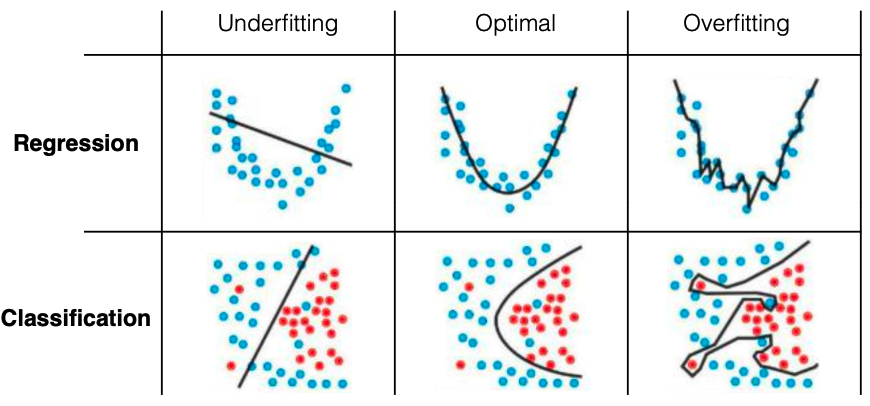
We have a finite number of samples $D = \{\mbf{x}_i,y_i\}_{i=1}^n$ that are generated from a function $f$ plus error so $y = f(\mbf{x}) + \epsilon$
Then we can decompose its error:
$$ \begin{aligned} \operatorname{MSE}\left(\hat{f}_{n}\right) &=\mathbb{E}_{\epsilon}\left[\left(y_{*}-\hat{f}_{n}\left(x_{*}\right)\right)^{2}\right] \\ &=\mathbb{E}\left[\left(\underbrace{\epsilon+f\left(x_{*}\right)}_{y_{*}}-\hat{f}_{n}\left(x_{*}\right)\right)^{2}\right] \end{aligned} $$Assume that there is a unknown and complex generator $\mathcal{D}$ that provides output pairs $(\mathbf{x},y)$.
In practice, in a real-world problem no one has access to $\mathcal{D}$ because problems are too complex
Try to write a computer program to generate all possible natural images that you can find in the word. Is it easy?
Let's assume here that we have access to $\mathcal{D}$ as a python function get_prob_under_D(x,y) that takes as input a pair (x,y) and returns the probability of the pair under $\mathcal{D}$.
If so, we can define the Bayes optimal classifier as the classifier that:
get_prob_under_D(x,y)
Bias and Variance terms go to zero!
$$\operatorname{MSE}\left(\hat{f}_{n}\right) = \underbrace{\tau^{2}}_{\text {Irreducible error }}$$
In order to accomplish a), let’s assume we have a loss or cost function:
$$\mathcal{L}(\underbrace{\hat{y}}_{pred.}, \underbrace{y}_{gt})\quad \text{where}\quad \hat{y}=h_{\theta}(\mathbf{x}) $$Ok we train on train split and we test on test split.
How do we select the distance and $k$ neighbours?
Ok we train on train split and we test on test split.
How do we select the distance and $k$ neighbours?
| Train | 70% | 60% | 80% |
|---|---|---|---|
| Validation (dev) | 20% | 20% | 10% |
| Test | 10% | 20% | 10% |
However, by partitioning the available data into three sets, we drastically reduce the number of samples which can be used for learning the model, and the results can depend on a particular random choice for the pair of (train, validation) sets.
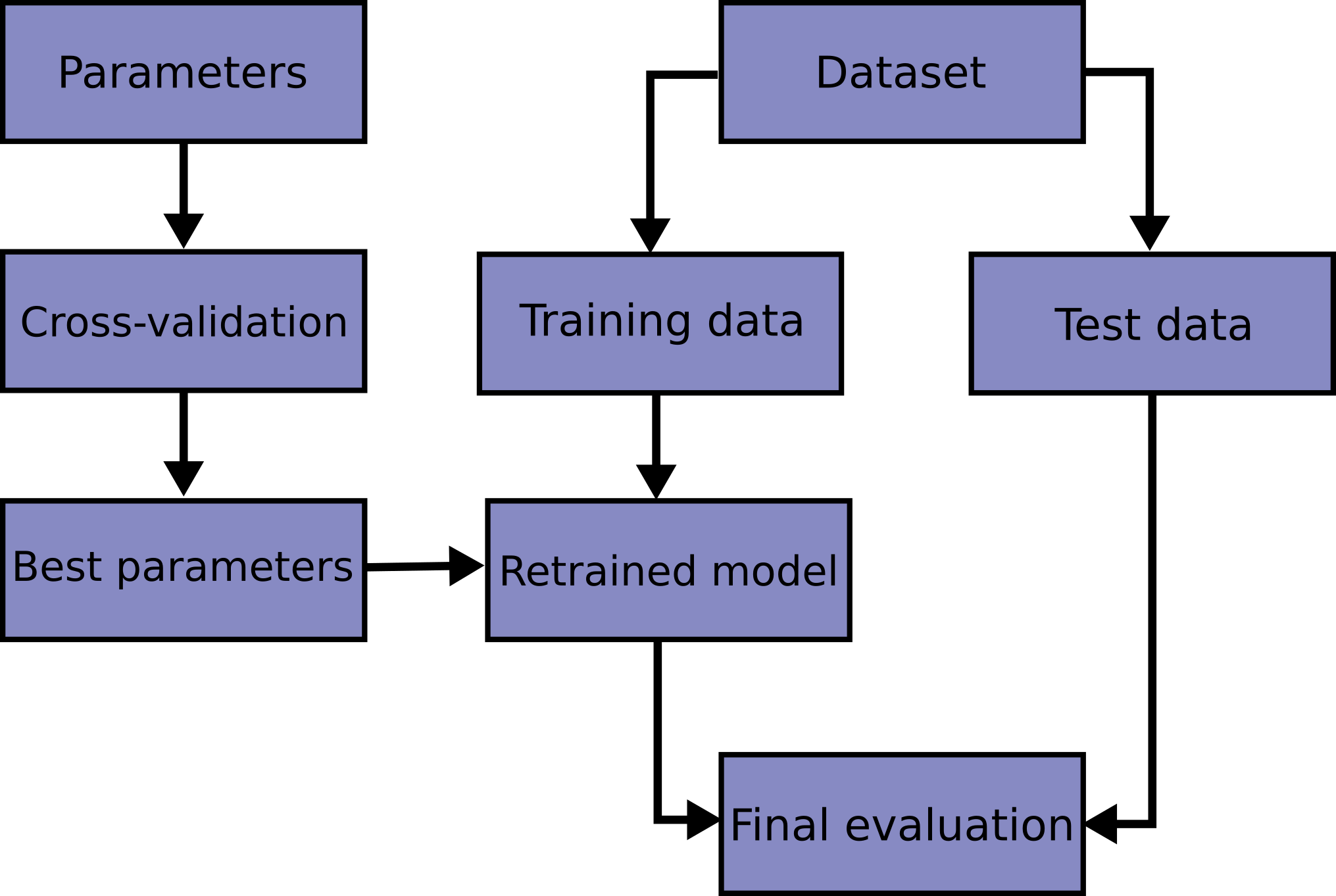
A solution to this problem is a procedure called cross-validation (CV for short).
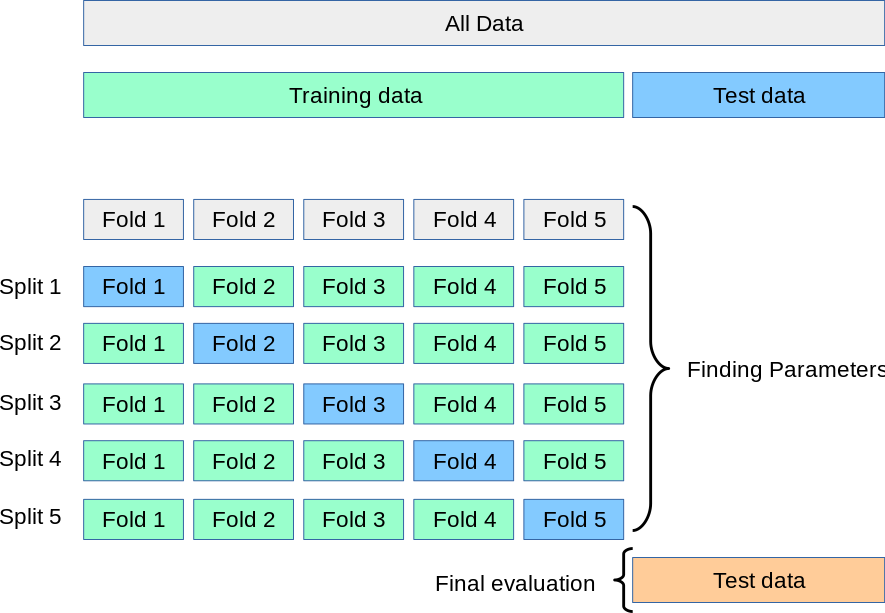
Pro:
Con:

In general:
Rule of thumb:
K are 2, 5, 10. 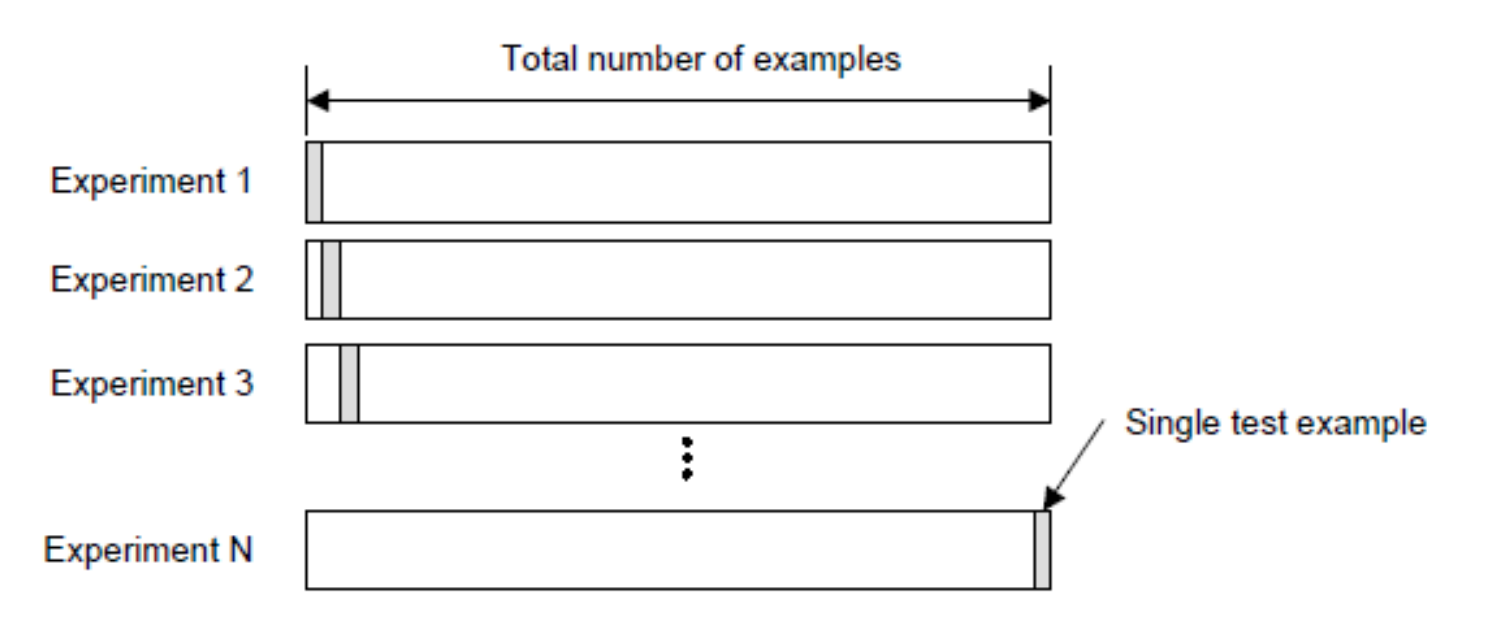
LOO is more computationally expensive than K-fold cross validation (not so expensive for K-NN).
In terms of accuracy, LOO often results in high variance as an estimator for the test error.
Since the samples are used to build each model, models constructed from folds are virtually identical to each other and to the model built from the entire training set.
As a general rule, most authors, and empirical evidence, suggest that 5- or 10- fold cross validation should be preferred to LOO.
Note sklearn provides interface for this but is mandatory to have in your mind the entire flow:
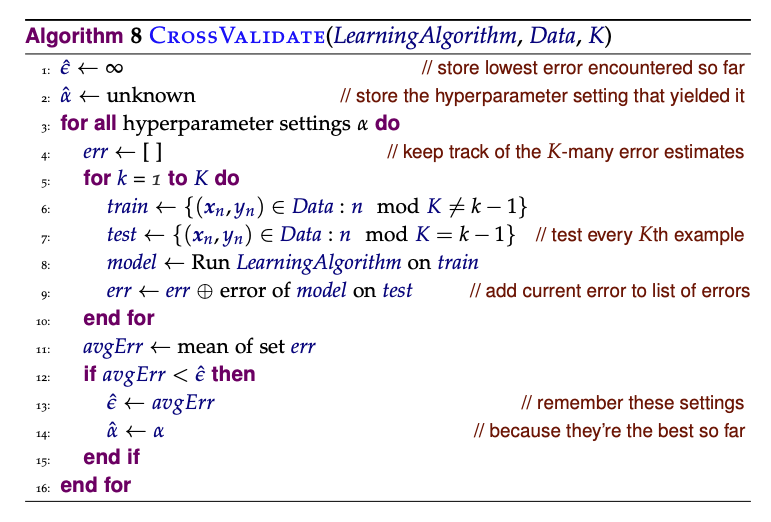

In this case, it is important that when you cross validate (or use development data), you do so over images, not over pixels.
The same goes for text problems where you sometimes want to classify things at a word level, but are handed a collection of documents. The important thing to keep in mind is that it is the images (or documents) that are drawn independently from your data distribution and not the pixels (or words), which are drawn dependently.
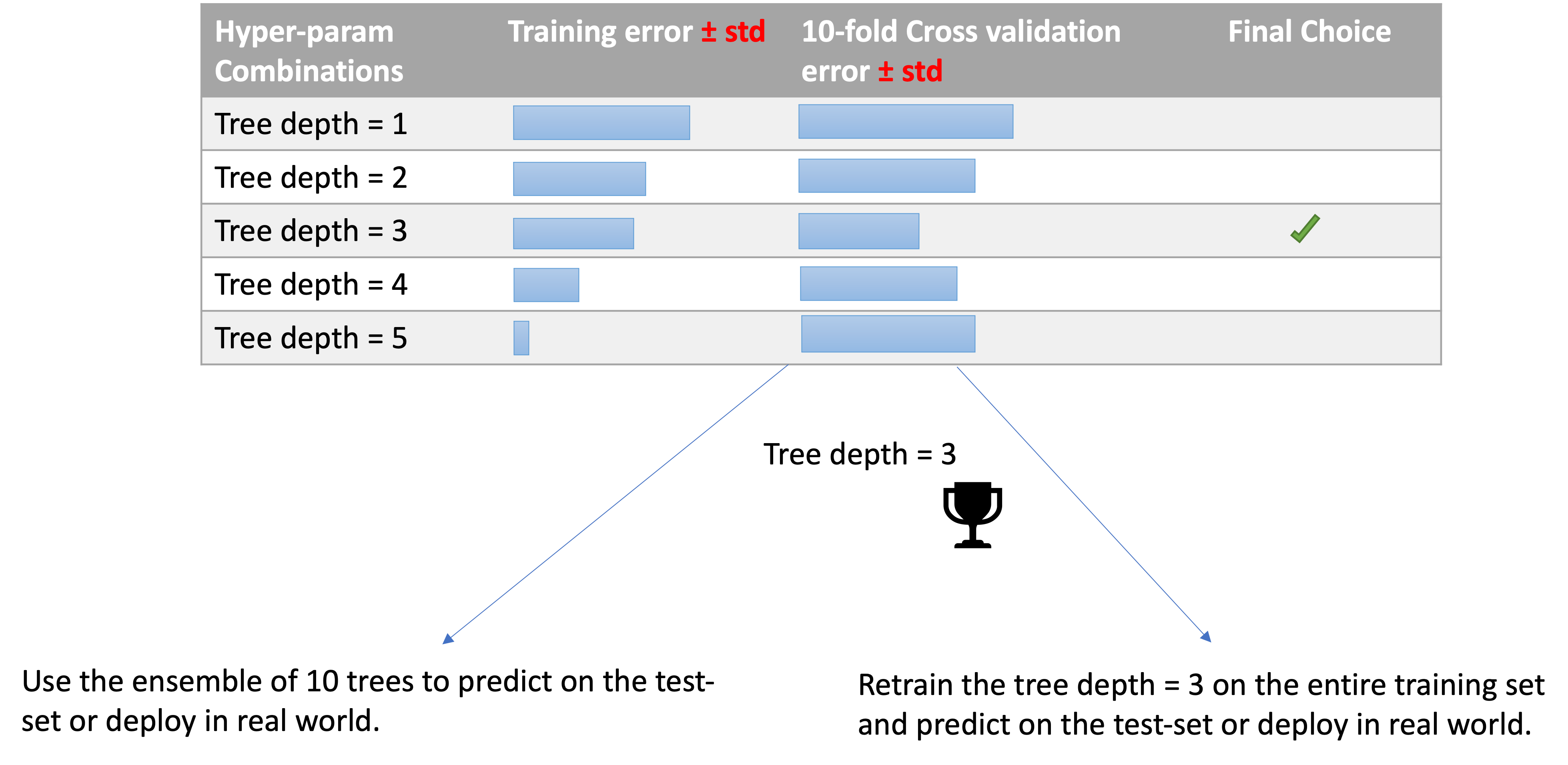
After running cross validation, you have 2 choices.
2. is generally preferred to 1.

We are working in the medical sector and we are using decision tree for their interpretability power, but we have to decide the depth of the tree.
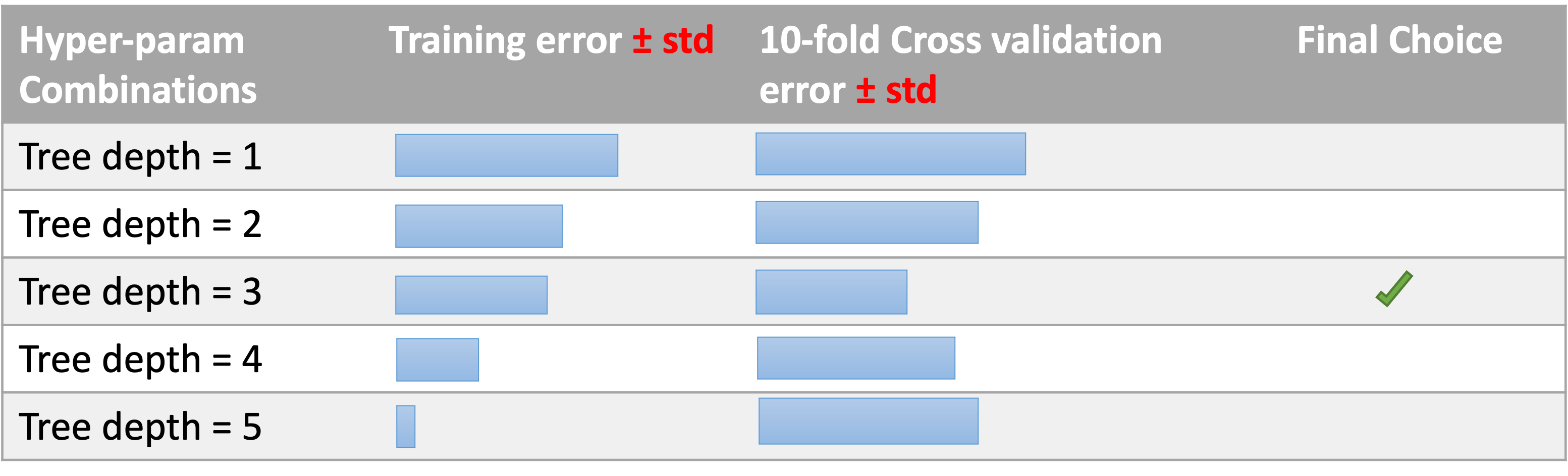

Exam look-alike question: how many models (decision trees here) you need to train to make the choice?
CV assumptions:
⚠️

Ensures that the same group is NOT represented in both testing and training sets. For example if the data is obtained from different subjects with several samples per-subject and if the model is flexible enough to learn from highly person specific features it could fail to generalize to new subjects.
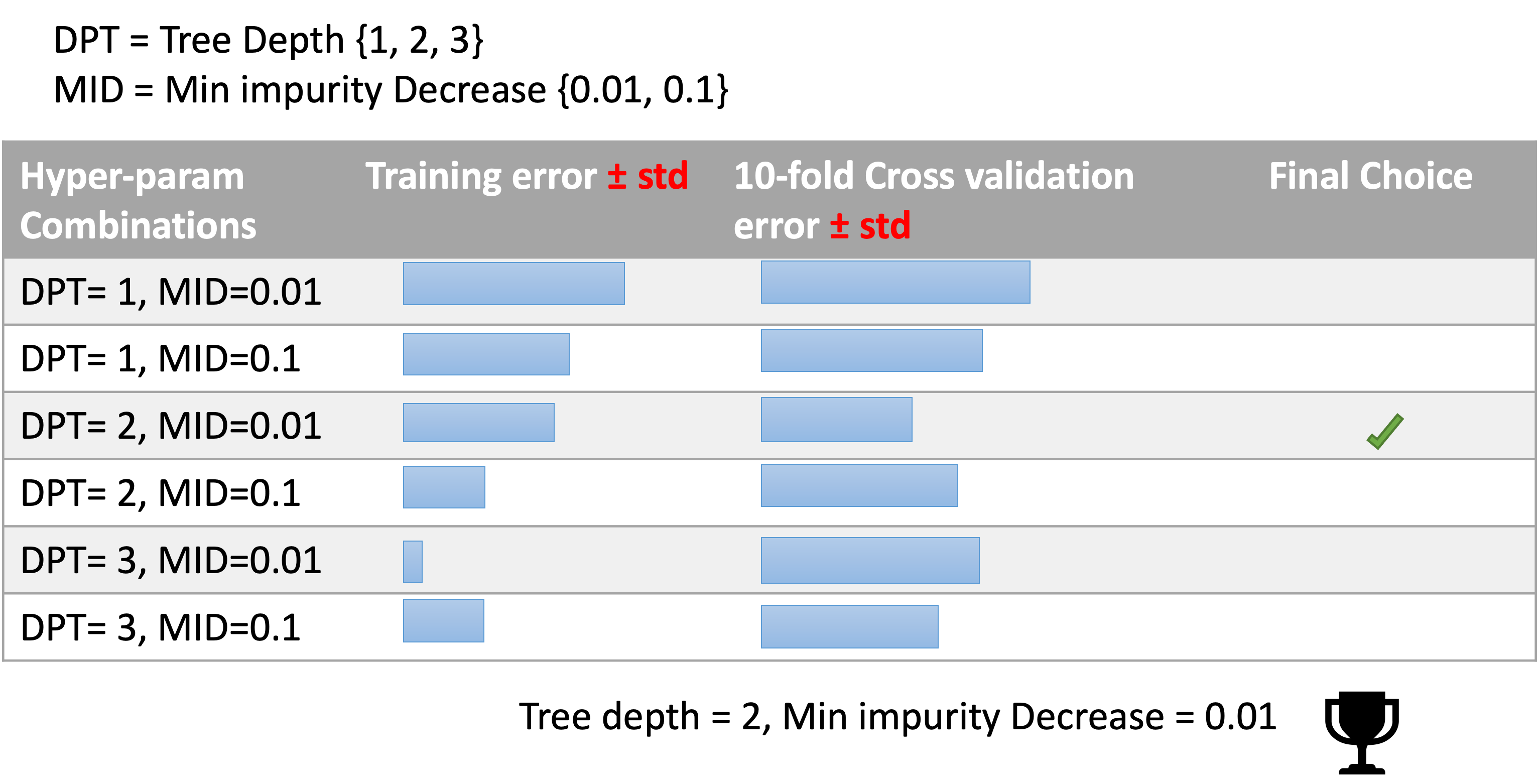
A search of HP consists of:
We are working in the medical sector and we are using decision trees for their interpretability power, but we have to decide the depth of the tree.
max_depth=None, # we know the best for this is 3 but what about the other?
min_impurity_decrease=0.0
max_depth=None, # we know the best for this is 3 but what about the other?
min_impurity_decrease=0.0

max_depth=None,
min_impurity_decrease=0.0
min_samples_leaf=1,
max_leaf_nodes=None,
# Set the parameters by cross-validation
tuned_parameters = [
{"max_depth": [1, 2, 3, None],
"min_impurity_decrease": [0.001, 0.01, 0.1, ],
"criterion": ["gini", "entropy"],
}
]
....
clf = GridSearchCV(DTC(), tuned_parameters, scoring="%s" % score)
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.tree import DecisionTreeClassifier as DTC
# Loading the Digits dataset
digits = datasets.load_digits()
# To apply a classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
X = digits.images.reshape((n_samples, -1))
y = digits.target
# Split the dataset in two parts
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=0)
# Set the parameters by cross-validation
tuned_parameters = [
{"max_depth": [1, 2, 3, None],
"min_impurity_decrease": [0.001, 0.01, 0.1, ],
"criterion": ["gini", "entropy"],
}
]
scores = ["accuracy"]
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
clf = GridSearchCV(DTC(), tuned_parameters, scoring="%s" % score)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_["mean_test_score"]
stds = clf.cv_results_["std_test_score"]
for mean, std, params in zip(means, stds, clf.cv_results_["params"]):
print("%0.3f (+/-%0.03f) for %r" % (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# Note the problem is too easy: the hyperparameter plateau is too flat and the
# output model is the same for precision and recall with ties in quality.
# Tuning hyper-parameters for accuracy
Best parameters set found on development set:
{'criterion': 'entropy', 'max_depth': None, 'min_impurity_decrease': 0.001}
Grid scores on development set:
0.209 (+/-0.007) for {'criterion': 'gini', 'max_depth': 1, 'min_impurity_decrease': 0.001}
0.209 (+/-0.007) for {'criterion': 'gini', 'max_depth': 1, 'min_impurity_decrease': 0.01}
0.107 (+/-0.003) for {'criterion': 'gini', 'max_depth': 1, 'min_impurity_decrease': 0.1}
0.328 (+/-0.007) for {'criterion': 'gini', 'max_depth': 2, 'min_impurity_decrease': 0.001}
0.328 (+/-0.007) for {'criterion': 'gini', 'max_depth': 2, 'min_impurity_decrease': 0.01}
0.107 (+/-0.003) for {'criterion': 'gini', 'max_depth': 2, 'min_impurity_decrease': 0.1}
0.479 (+/-0.026) for {'criterion': 'gini', 'max_depth': 3, 'min_impurity_decrease': 0.001}
0.478 (+/-0.024) for {'criterion': 'gini', 'max_depth': 3, 'min_impurity_decrease': 0.01}
0.107 (+/-0.003) for {'criterion': 'gini', 'max_depth': 3, 'min_impurity_decrease': 0.1}
0.842 (+/-0.045) for {'criterion': 'gini', 'max_depth': None, 'min_impurity_decrease': 0.001}
0.786 (+/-0.071) for {'criterion': 'gini', 'max_depth': None, 'min_impurity_decrease': 0.01}
0.107 (+/-0.003) for {'criterion': 'gini', 'max_depth': None, 'min_impurity_decrease': 0.1}
0.200 (+/-0.013) for {'criterion': 'entropy', 'max_depth': 1, 'min_impurity_decrease': 0.001}
0.200 (+/-0.013) for {'criterion': 'entropy', 'max_depth': 1, 'min_impurity_decrease': 0.01}
0.200 (+/-0.013) for {'criterion': 'entropy', 'max_depth': 1, 'min_impurity_decrease': 0.1}
0.351 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 2, 'min_impurity_decrease': 0.001}
0.351 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 2, 'min_impurity_decrease': 0.01}
0.351 (+/-0.025) for {'criterion': 'entropy', 'max_depth': 2, 'min_impurity_decrease': 0.1}
0.527 (+/-0.039) for {'criterion': 'entropy', 'max_depth': 3, 'min_impurity_decrease': 0.001}
0.527 (+/-0.039) for {'criterion': 'entropy', 'max_depth': 3, 'min_impurity_decrease': 0.01}
0.519 (+/-0.050) for {'criterion': 'entropy', 'max_depth': 3, 'min_impurity_decrease': 0.1}
0.845 (+/-0.025) for {'criterion': 'entropy', 'max_depth': None, 'min_impurity_decrease': 0.001}
0.827 (+/-0.041) for {'criterion': 'entropy', 'max_depth': None, 'min_impurity_decrease': 0.01}
0.614 (+/-0.057) for {'criterion': 'entropy', 'max_depth': None, 'min_impurity_decrease': 0.1}
Detailed classification report:
The model is trained on the full development set.
The scores are computed on the full evaluation set.
precision recall f1-score support
0 0.92 0.89 0.91 27
1 0.88 0.86 0.87 35
2 0.93 0.78 0.85 36
3 0.69 0.93 0.79 29
4 0.86 0.83 0.85 30
5 0.87 0.82 0.85 40
6 0.97 0.86 0.92 44
7 0.90 0.97 0.94 39
8 0.78 0.90 0.83 39
9 0.82 0.76 0.78 41
accuracy 0.86 360
macro avg 0.86 0.86 0.86 360
weighted avg 0.87 0.86 0.86 360
A budget can be chosen independent of the number of parameters and possible values.
Adding parameters that do not influence the performance does not decrease efficiency.
{'C': scipy.stats.expon(scale=100), 'gamma': scipy.stats.expon(scale=.1),
'kernel': ['rbf'], 'class_weight':['balanced', None]}
n_iter int, default=10
Number of parameter settings that are sampled. n_iter trades off runtime vs quality of the solution.
Idea: Successive halving (SH) is like a tournament among candidate parameter combinations.
number of training samples, but it can also be an arbitrary numeric parameter such as n_estimators in a random forest.| Person ID (training example) |
Overcooked pasta? | Waiting Time | Rude Waiter? | Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
How to approach this, First, Analysis:
| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
So the starting set if $\vert S \vert = 5$, and $H(S)=-\sum_{y \in \{yes,no\}} p(y)\log_2(p(y))$
Over $S$, $p(y==yes) = \frac{3}{5}$ while $p(y==no) = \frac{2}{5}$
So, $H(S)= -\frac{3}{5}\log_2(\frac{3}{5})-\frac{2}{5}\log_2(\frac{2}{5}) \approx 0.971$
| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
$\frac{3}{5}H(y~|~\texttt{Ov. pasta==Yes}) = ~~\texttt{?}$
| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
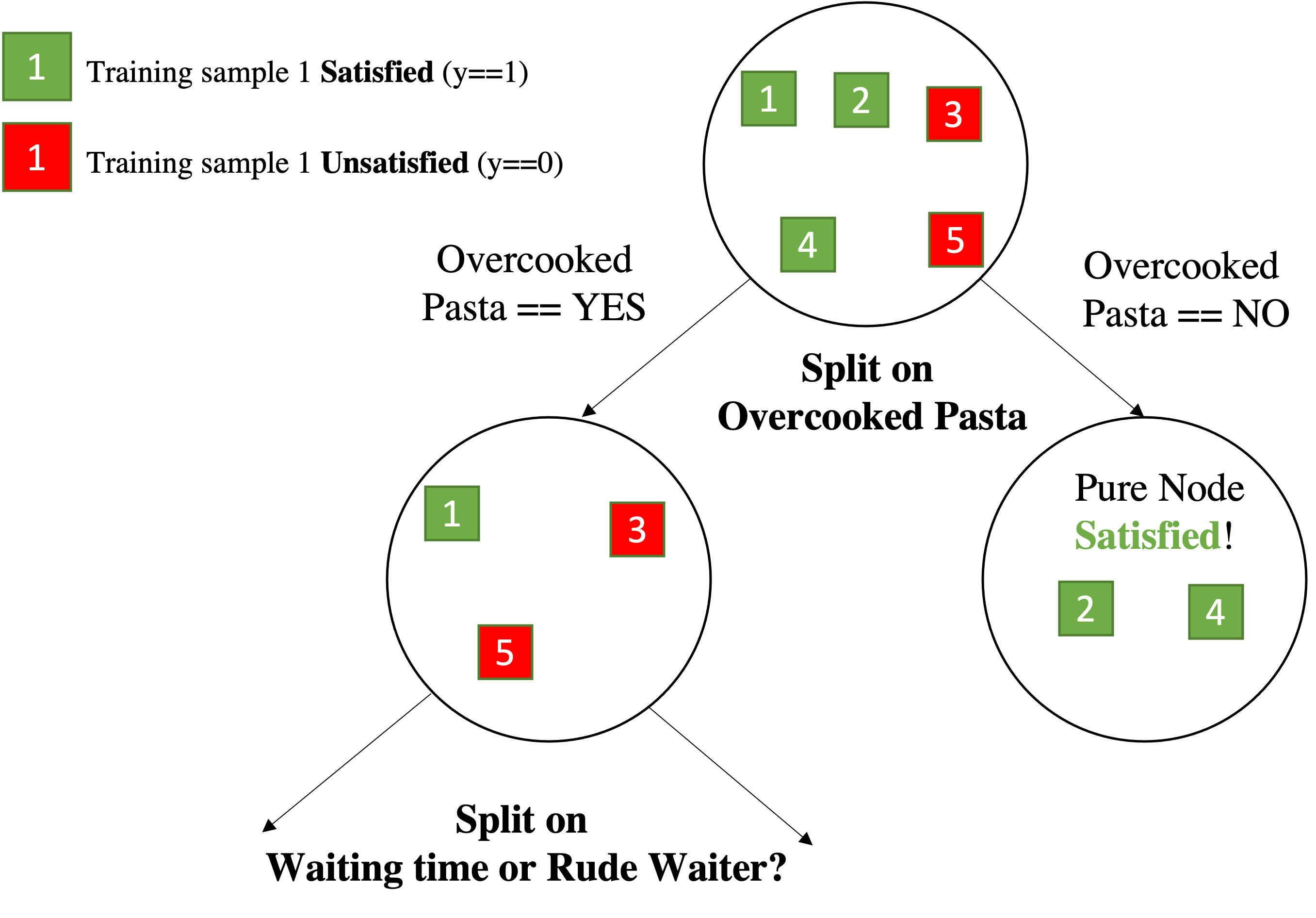
Split on Overcooked pasta and compute the new Entropy. I have two case:
Now look at the labels for $\texttt{Ov. pasta==Yes}$ $$\frac{3}{5}H(y~|~\texttt{Ov. pasta==Yes}) = \frac{3}{5} \cdot \biggl[-\sum_{y \in \{yes,no\}} p(y|x)\log_2(p(y|x))\biggl]$$
we have 2 No Satisfaction and 1 Yes,Satisfaction , over 3 samples after choosing:
$$\frac{3}{5}H(y~|~\texttt{Ov. pasta==Yes}) =\frac{3}{5}\cdot \big[ \underbrace{-\frac{1}{3}\log_2(\frac{1}{3})}_{yes}-\underbrace{\frac{2}{3}\log_2(\frac{2}{3})}_{no}\big] \approx 0.551$$| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
Split on Overcooked pasta and compute the new Entropy. I have two case:
Now look at the labels for $\texttt{Ov. pasta==No}$ $$\frac{2}{5}H(y~|~\texttt{Ov. pasta==No}) = \frac{2}{5} \cdot \biggl[-\sum_{y \in \{yes,no\}} p(y|x)\log_2(p(y|x))\biggl]$$
we have 0 UnSatisfaction and 2 Yes, Satisfaction , over 2 samples after choosing:
$$\frac{2}{5}H(y~|~\texttt{Ov. pasta==No}) =\frac{2}{5}\cdot \big[ -\frac{0}{2}\log_2(\frac{0}{2})-\frac{2}{2}\log_2(\frac{2}{2}) \big] = 0$$| Person ID (training example) |
[Feat. 1] Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Yes | Long | No | 1 (yes) |
| $\mathbf{x}_2$ | No | Short | Yes | 1 (yes) |
| $\mathbf{x}_3$ | Yes | Long | Yes | 0 (no) |
| $\mathbf{x}_4$ | No | Long | Yes | 1 (yes) |
| $\mathbf{x}_5$ | Yes | Short | Yes | 0 (no) |
Information Gain for split on $\texttt{Overcooked pasta}$:
$$IG(Y~|~\texttt{Ov. pasta}) \doteq \frac{5}{5}H(S) - \Big[ \frac{2}{5}H(y~|~\texttt{Ov. pasta==No}) + \frac{3}{5}H(y~|~\texttt{Ov. pasta==Yes}) \Big] $$$$IG(Y~|~\texttt{Ov. pasta}) \doteq 0.971 - [0.551+0] \approx 0.42 $$| Split Feature | IG |
|---|---|
| Overcooked Pasta | 0.42 |
| Waiting Time | 0.020 |
| Rude Waiter | 0.171 |
So we split on Overcooked Pasta over Yes with a gain of $0.42$
Overcooked Pasta is a binary variable so splitting over Yes or Not does not matter.Overcooked Pasta were $\{Yes, No, Maybe\}$ we should have splitted taking note of the threshold.
-i.e. if we binary split on Yes, the other split is $\neg$ Yes (which means it could be No or Maybe).| Person ID (training example) |
Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_1$ | Long | No | 1 (yes) | |
| $\mathbf{x}_3$ | Long | Yes | 0 (no) | |
| $\mathbf{x}_5$ | Short | Yes | 0 (no) |
| Person ID (training example) |
Overcooked pasta? |
[Feat. 2] Waiting Time |
[Feat. 3] Rude Waiter? |
Satisfied $y$ |
|---|---|---|---|---|
| $\mathbf{x}_2$ | Short | Yes | 1 (yes) | |
| $\mathbf{x}_4$ | Long | Yes | 1 (yes) |

from sklearn.metrics import confusion_matrix
y_true = [2, 0, 2, 2, 0, 1]
y_pred = [0, 0, 2, 2, 0, 2]
confusion_matrix(y_true, y_pred)
> array([[2, 0, 0],
[0, 0, 1],
[1, 0, 2]])
{{import matplotlib.pyplot as plt; from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay;y_true = [2, 0, 2, 2, 0, 1];y_pred = [0, 0, 2, 2, 0, 2];disp = ConfusionMatrixDisplay(confusion_matrix(y_true, y_pred));disp.plot();plt.show()}}
Ability to return only images that match the query
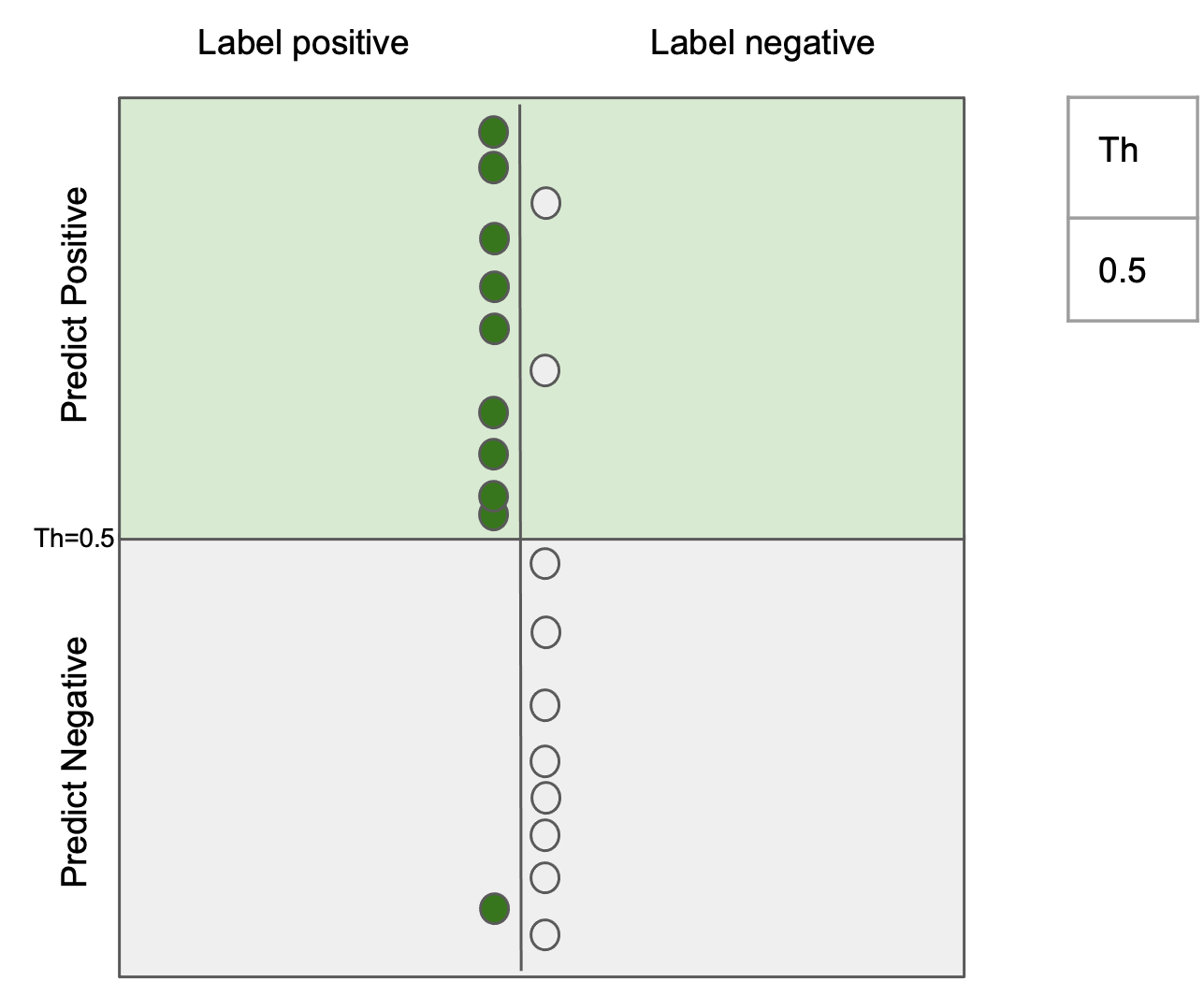
$$ \text{precision} = \frac{tp}{tp + fp}$$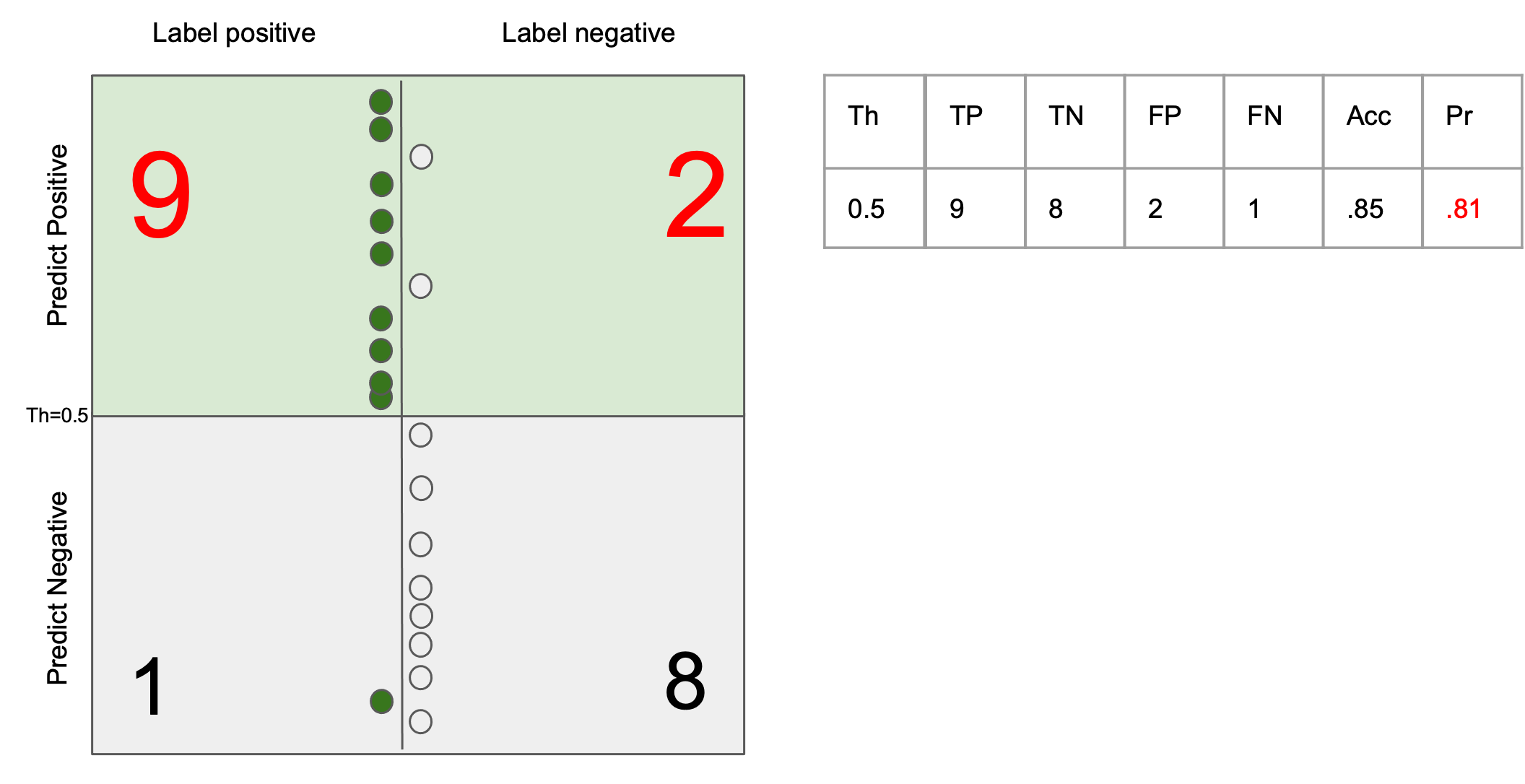
Ability to return all the images that match the query
$$ \text{recall} = \frac{tp}{tp + fn} $$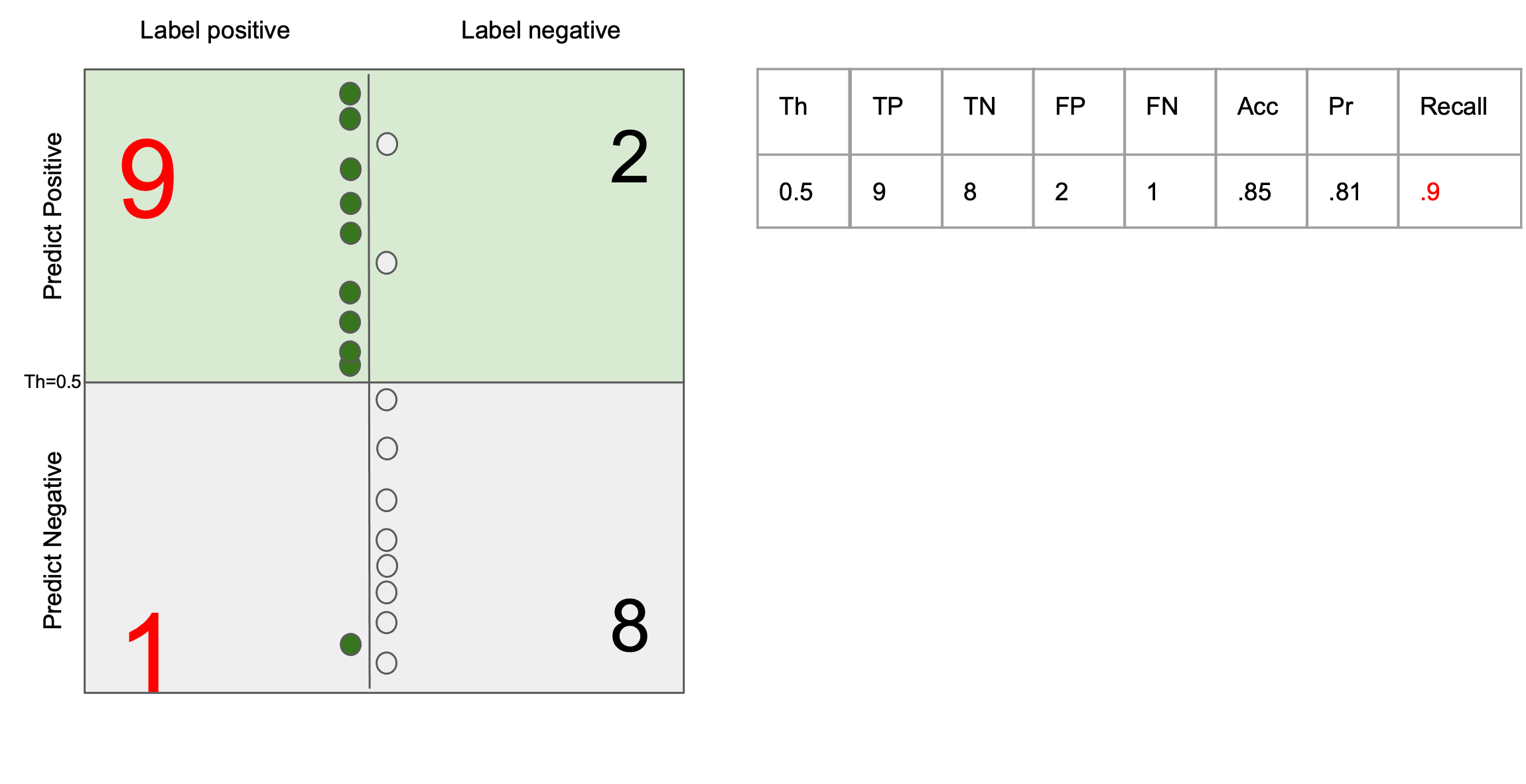
y = np.array([0, 0, 1, 1, 1, 0])
scores = np.array([0.1, 0.4, 0.35, 0.8, 0.01, 0.2])
import numpy as np
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y, scores, drop_intermediate=False)
plt.figure()
lw = 2
plt.plot(
fpr,
tpr,
color="darkorange",
lw=lw,
label="ROC curve (area = %0.2f)" % auc(fpr, tpr),
)
plt.plot([0, 1], [0, 1], color="navy", lw=lw, linestyle="--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver operating characteristic example")
plt.legend(loc="lower right")
plt.show()
def thrs_count(scores, index, denom, thrs, normalize=True):
count = sum(scores[index] >= thrs)
return count/denom if normalize else count
pos = y == 1 # indexing the positive
neg = y == 0 # indexing the negative
n_pos = sum(pos) # sum the ground-truth positve
n_neg = sum(neg) # sum the ground-truth negative
sort_scores = sorted(scores, reverse=True) # sort highest first
sort_scores = [sort_scores[0] + 1] + sort_scores # add extra point
ROC = [(thrs_count(scores, pos, n_pos, thrs), thrs_count(scores, neg, n_neg, thrs))
for thrs in sort_scores] # for all thresholds, count TP and FP and normalize them
my_TPR, my_FPR = list(map(lambda *args: args, *ROC)) # reshape
assert all([np.allclose(tpr, my_TPR),
np.allclose(fpr, my_FPR),
np.allclose(thresholds, sort_scores)]), 'Your ROC is wrong'
plt.figure(figsize=(7,7))
plt.plot(my_FPR,my_TPR);
plt.plot([0, 1], [0, 1], color="navy", lw=lw, linestyle="--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver operating characteristic example")
plt.show()
AUC = np.dot(np.diff(fpr), tpr[1:])
print(AUC)
assert np.allclose(AUC, auc(fpr, tpr)), 'my AUC is WRONG'
0.5555555555555556
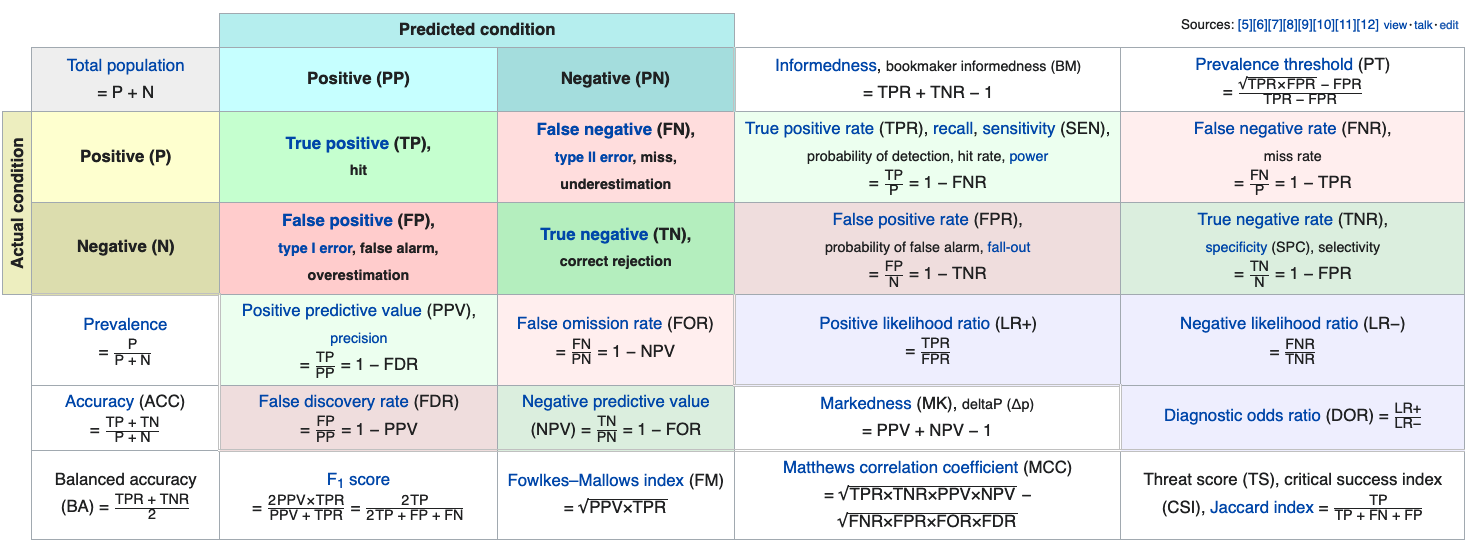
$ \text{precision} = \frac{tp}{tp + fp}$
$ \text{recall} = \frac{tp}{tp + fn} $
$F_\beta = (1 + \beta^2) \frac{\text{precision} \times \text{recall}}{\beta^2 \text{precision} + \text{recall}} $
Symptom: Prevalence < 5% (no strict definition) $\frac{P_{gt}}{P_{gt}+N_{gt}}$
In general: Accuracy << AUROC << AUPRC
Avoids inflated performance estimates on imbalanced datasets. It is the raw accuracy where each sample is weighted according to the inverse prevalence of its true class.
Thus for balanced datasets, the score is equal to accuracy.
In the binary case, balanced accuracy is equal to the arithmetic mean of sensitivity (true positive rate) and specificity (true negative rate):
$$ \texttt{balanced-accuracy} = \frac{1}{2}\left( \frac{TP}{TP + FN} + \frac{TN}{TN + FP}\right )$$You are given a machine learning system performing binary classification on $6$ samples. The ground-truth labels and the unnormalized scores for the six samples are:
| labels | -1 | 1 | -1 | 1 | -1 | 1 |
|---|---|---|---|---|---|---|
| score | 1000 | 0.5 | -0.9 | 0.8 | -0.1 | 0.1 |
The scores are unnormalized (i.e. not probabilities) and correlated with the positive label (1).
| labels | -1 | 1 | -1 | 1 | -1 | 1 |
|---|---|---|---|---|---|---|
| score | ? | ? | ? | ? | ? | ? |
| labels | -1 | 1 | -1 | 1 | -1 | 1 |
|---|---|---|---|---|---|---|
| score | ? | ? | ? | ? | ? | ? |
y = np.array([-1, 1, -1, 1, -1, 1])
scores = np.array([1000, 0.5, -0.9, 0.8, -0.1, 0.1])
import numpy as np
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y, scores, drop_intermediate=False, pos_label=1)
plt.figure()
lw = 2
plt.plot(
fpr,
tpr,
color="darkorange",
lw=lw,
label="ROC curve (area = %0.2f)" % auc(fpr, tpr),
)
plt.plot([0, 1], [0, 1], color="navy", lw=lw, linestyle="--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver operating characteristic example")
plt.legend(loc="lower right")
plt.show()
pos = y == 1 # indexing the positive
neg = y == -1 # indexing the negative
n_pos = sum(pos) # sum the ground-truth positve
n_neg = sum(neg) # sum the ground-truth negative
sort_scores = sorted(scores, reverse=True) # sort highest first
sort_scores = [sort_scores[0] + 1] + sort_scores # add extra point
ROC = [(thrs_count(scores, pos, n_pos, thrs, normalize=False),
thrs_count(scores, neg, n_neg, thrs, normalize=False))
for thrs in sort_scores] # for all thresholds, count TP and FP and normalize them
diff_fpr = np.diff(np.array(my_FPR))
my_TPR, my_FPR = list(map(lambda *args: args, *ROC)) # reshape
table = '|thrs |tpr |fpr |diff_fpr |\n|--- |--- |--- |--- \n'
for count, (mtpr, mfpr, thrs) in enumerate(zip(my_TPR, my_FPR, sort_scores)):
diff_fpr = mfpr/n_neg-my_FPR[count-1]/n_neg
table += f'|{thrs} |{mtpr}/{n_pos} |{mfpr}/{n_neg} | { str(diff_fpr)[:5] if count > 0 else 0} \n'
print(table)
|thrs |tpr |fpr |diff_fpr | |--- |--- |--- |--- |1001.0 |0/3 |0/3 | 0 |1000.0 |0/3 |1/3 | 0.333 |0.8 |1/3 |1/3 | 0.0 |0.5 |2/3 |1/3 | 0.0 |0.1 |3/3 |1/3 | 0.0 |-0.1 |3/3 |2/3 | 0.333 |-0.9 |3/3 |3/3 | 0.333
NameError: name 'table' is not defined
y = np.array([-1, 1, -1, 1, -1, 1])
alpha = 1
scores = np.array([-alpha, alpha, -alpha, alpha, -alpha, alpha])
import numpy as np
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y, scores, drop_intermediate=False, pos_label=1)
plt.figure()
lw = 2
plt.plot(
fpr,
tpr,
color="darkorange",
lw=lw,
label="ROC curve (area = %0.2f)" % auc(fpr, tpr),
)
plt.plot([0, 1], [0, 1], color="navy", lw=lw, linestyle="--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver operating characteristic example")
plt.legend(loc="lower right")
plt.show()
y = np.array([-1, 1, -1, 1, -1, 1])
alpha = -1
scores = np.array([-alpha, alpha, -alpha, alpha, -alpha, alpha])
import numpy as np
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y, scores, drop_intermediate=False, pos_label=1)
plt.figure()
lw = 2
plt.plot(
fpr,
tpr,
color="darkorange",
lw=lw,
label="ROC curve (area = %0.2f)" % auc(fpr, tpr),
)
plt.plot([0, 1], [0, 1], color="navy", lw=lw, linestyle="--")
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.title("Receiver operating characteristic example")
plt.legend(loc="lower right")
plt.show()