import numpy as np
from matplotlib import pyplot as plt
from sklearn import linear_model, datasets
n_samples = 100
size = 10
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=2,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot(projection='3d')
ax.scatter(X[..., 0], X[..., 1], y, c='red', marker='o')
ax.view_init(0, -90)
import numpy as np
from matplotlib import pyplot as plt
from sklearn import linear_model, datasets
n_samples = 100
size = 10
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=2,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot(projection='3d')
ax.scatter(X[..., 0], X[..., 1], y, c='red', marker='o')
ax.view_init(0, -90)
| x_1 | x_2 | y |
|---|---|---|
| -1.415 | -0.420 | -116.6 |
| 0.5219 | 0.2969 | 58.737 |
| -0.889 | -0.815 | -73.89 |
| -0.883 | 0.1537 | -113.3 |
| 0.7384 | 0.1713 | 63.998 |
| -0.264 | 2.7201 | -30.33 |
| 1.1428 | 0.7519 | 81.616 |
| 0.3613 | 1.5380 | 48.283 |
| 0.8125 | 1.3562 | 119.06 |
| -0.223 | 0.7140 | -12.36 |
| -1.106 | -1.196 | -122.4 |
We want to regress $y$ from $\mbf{x}$ that is we want to learn a function $f_{\boldsymbol{\theta}}$ parametrized by some parameters $\boldsymbol{\theta}$ so that $ f_{\boldsymbol{\theta}}: \underbrace{\mbf{x}}_{\text{input}} \mapsto \underbrace{y}_{\text{output}}$
We assume the relation $f \longleftrightarrow y$ is linear.
We know $D = \{\mbf{x}_i,y_i\}_{i=1}^n$ and we want to find ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d)$
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \theta_0+ \theta_1\cdot x_1 + \theta_2\cdot x_2 + \ldots + \theta_d\cdot x_d$$So ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d) \in \mathbb{R}^{d+1}$.
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \big(\sum_{i=1}^d\theta_i\cdot x_i\big) + \theta_0$$We can augment each feature to have a bias (intercept term) set to $1$ so that $\mbf{x} \doteq [1,\mbf{x}]$.
Doing so $\mbf{x} \in \mathbb{R}^{d+1}$
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \theta_0\cdot \underbrace{x_0}_{\text{always}~1} + \theta_1\cdot x_1 + \theta_2\cdot x_2 + \ldots + \theta_d\cdot x_d$$So ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d) \in \mathbb{R}^{d+1}$.
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^d\theta_i\cdot x_i = \bmf{\theta}^T\mbf{x} $$No matter how many training points $N$ you have, the parameters are fixed in $\bmf{\theta}$.
Note that $\bmf{\theta} \in \mathbb{R}^{d+1}$.
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^d\theta_i\cdot x_i = \bmf{\theta}^T\mbf{x} $$You see now that the loss is more explicit compared to non-parametric models (K-NN, Decision Trees).
$$ \mathcal{J}(\mbf{\theta};\mbf{x},y)= \frac{1}{2} \sum_{i=1}^{n}\mathcal{L}\big(y_{i}, f_{\boldsymbol{\theta}}(\mbf{x}_i)\big)$$where
$$\mathcal{L}\big(y, f_{\boldsymbol{\theta}}(\mbf{x})\big) = \big(f_{\boldsymbol{\theta}}(\mbf{x}) - y \big)^2 $$The loss is the squared error.
We need to minimize $$ \mathcal{J}(\mbf{\theta};\mbf{x},y)= \frac{1}{2} \sum_{i=1}^{n}\mathcal{L}\big(y_{i}, f_{\boldsymbol{\theta}}(\mbf{x}_i)\big)$$ so to find:
$$\mbf{\theta}^{\star} = \arg\min_{\mbf{\theta}} \mathcal{J}(\mbf{\theta};\mbf{x},y) $$We define the design matrix $\mbf{X} \in \mathbb{R}^{n\times (d+1)}$ and label matrix $\mbf{y}\in \mathbb{R}^{n\times 1} $
$$ \def\horzbar{\rule[.5ex]{2.5ex}{0.5pt}} \mbf{X} = \left[ \begin{array}{ccc} \horzbar & \mbf{x}^{T}_{1} & \horzbar \\ \horzbar & \mbf{x}^{T}_{2} & \horzbar \\ & \vdots & \\ \horzbar & \mbf{x}^{T}_{n} & \horzbar \end{array} \right] \; \qquad \mbf{y} = \left[ \begin{array}{c} {y}_{1} \\ {y}_{2} \\ \vdots \\ {y}_{n} \\ \end{array} \right] $$The parameters $\bmf{\theta} \in \mathbb{R}^{d+1}$ are:
$$ \bmf{\theta} = \left[ \begin{array}{c} \theta_{0} \\ \theta_{1} \\ \vdots \\ \theta_{d} \\ \end{array} \right] $$Set the gradient to zero to find critical points:
$$\nabla_{\mbf{\theta}} \mathcal{J}(\mbf{\theta};\mbf{X},\mbf{y}) = 0$$For now forget the "equal to zero":
$$\nabla_{\mbf{\theta}} \frac{1}{2} \big[ (\mbf{X}\bmf{\theta})^T(\mbf{X}\bmf{\theta}) - \underbrace{(\mbf{X}\bmf{\theta})^T\mbf{y}}_{\text{scalar}} - \underbrace{\mbf{y}^T(\mbf{X}\bmf{\theta})}_{\text{scalar}} + \mbf{y}^T\mbf{y}) \big] $$Assumes $\mbf{X}^T\mbf{X}$ is invertible:
$$\bmf{\theta} = \underbrace{(\mbf{X}^T\mbf{X})^{-1}\mbf{X}^T}_{\text{pseudo inverse}}\mbf{y} = \mbf{X}^{+}\mbf{y}$$where:
$$ \mbf{X}^{+} \doteq (\mbf{X}^T\mbf{X})^{-1}\mbf{X}^T$$%matplotlib notebook
from sklearn import linear_model, datasets
from matplotlib import pyplot as plt
import numpy as np
n_samples = 100
size = 8
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=2,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot(projection='3d')
# Linear Regression
bias = np.ones((X.shape[0], 1))
X = np.hstack((X, bias))
theta = np.linalg.inv(X.T@X)@X.T@y
# Now MeshGrid
Xmin, Xmax = X.min(), X.max()
support = np.linspace(Xmin, Xmax, 10)
xx, yy = np.meshgrid(support, support)
data = np.stack((xx, yy), axis=2)
data = data.reshape(-1, 2)
data = np.hstack((data, np.ones((data.shape[0], 1))))
z = np.dot(theta, data.T)
z = z.reshape(xx.shape)
ax.plot_surface(xx, yy, z, alpha=0.2)
ax.scatter(X[..., 0], X[..., 1], y, c='red', marker='o')
ax.view_init(0, 90)
%matplotlib inline
plt.figure()
plt.bar(list(range(theta.size)),theta); plt.xlabel('Coefficients');
%matplotlib inline
plt.figure()
plt.bar(list(range(coef_gt.size)),coef_gt); plt.xlabel('Coefficients GT');
Assumes $\mbf{X}^T\mbf{X}$ is invertible (full rank) and $n\gg d$:
$$\bmf{\theta} = \underbrace{(\mbf{X}^T\mbf{X})^{-1}}_{d\times d}\underbrace{\mbf{X}^T}_{d\times n}\underbrace{\mbf{y}}_{n\times 1}$$it solves the linear system so that the plane best fits all the points with a trade-off given by the square of the residuals (least squares).
We can invert it "directly"
$$\bmf{\theta} = \mbf{X}^{-1}\mbf{y}$$Why? $$ (\mbf{A}\mbf{B})^{-1} = (\mbf{B}^{-1}\mbf{A}^{-1}) $$ then:
$$\bmf{\theta} = (\mbf{X}^T\mbf{X})^{-1}\mbf{X}^T\mbf{y} = \mbf{X}^{-1}(\mbf{X}^T)^{-1}\mbf{X}^T\mbf{y} = \mbf{X}^{-1}\mbf{y}$$%matplotlib notebook
from sklearn import linear_model, datasets
from matplotlib import pyplot as plt
import numpy as np
n_samples = 3
size = 12
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=2,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot(projection='3d')
ax.scatter(X[..., 0], X[..., 1], y, c='blue', marker='o')
# Linear Regression
bias = np.ones((X.shape[0], 1))
X = np.hstack((X, bias))
theta = np.linalg.inv(X)@y
# Now MeshGrid
Xmin, Xmax = X.min(), X.max()
support = np.linspace(Xmin, Xmax, 10)
xx, yy = np.meshgrid(support, support)
data = np.stack((xx, yy), axis=2)
data = data.reshape(-1, 2)
data = np.hstack((data, np.ones((data.shape[0], 1))))
z = np.dot(theta, data.T)
z = z.reshape(xx.shape)
ax.plot_surface(xx, yy, z, alpha=0.2)
ax.scatter(X[..., 0], X[..., 1], y, c='red', marker='o')
ax.view_init(0, 90)
We see the plane passes exactly through the "training points".
To go probabilistic, we have to make an assumption that each $y$ is generated linearly but with additive Gaussian Noise.
So
$$ y_i = \bmf{\theta}^T\mbf{x}_i + \epsilon$$where $$ \epsilon = \mathcal{N}(0,\sigma^2)$$
To go probabilistic, we have to make an assumption that each $y$ is generated linearly but with additive Gaussian Noise.
So
$$ \epsilon = y_i - \bmf{\theta}^T\mbf{x}_i \sim \mathcal{N}(0,\sigma^2)$$To go probabilistic, we have to make an assumption that each $y$ is generated linearly but with additive Gaussian Noise.
So
$$ \epsilon_i = y_i - \bmf{\theta}^T\mbf{x}_i \sim \mathcal{N}(0,\sigma^2)$$which means the errors behave IID from a Normal Distribution.
$$ p\left(\epsilon_i\right)=\frac{1}{\sqrt{2 \pi} \sigma} \exp \left(-\frac{\epsilon_i^{2}}{2 \sigma^{2}}\right) $$We look at the conditional probability of $y$ given $\mbf{x}$ aka $p(y|\mbf{x}; \bmf{\theta})$:
$$ p\left(y_i \mid x_i ; \theta\right)=\frac{1}{\sqrt{2 \pi} \sigma} \exp \left(-\frac{\left(y_i-\theta^{T} x_i\right)^{2}}{2 \sigma^{2}}\right) $$now is function of $y$ yet centered on $\theta^{T} x_i$: $$ y_i \mid x_i ; \theta \sim \mathcal{N}\left(\theta^{T} x_i, \sigma^{2}\right) $$
For a single training point:
$$ p\left(y_i \mid x_i ; \theta\right) \doteq L(\bmf{\theta};\mbf{x}_i,y_i) = \frac{1}{\sqrt{2 \pi} \sigma} \exp \left(-\frac{\left(y_i-\theta^{T} x_i\right)^{2}}{2 \sigma^{2}}\right) $$For multiple training point $\{\mbf{x}_i,y_i\}$, given IID assumptions on $\epsilon$ and thus $y|x$
$$ \begin{aligned} L(\theta) &=\prod_{i=1}^{n} p\left(y^{(i)} \mid x^{(i)} ; \theta\right) \\ &=\prod_{i=1}^{n} \frac{1}{\sqrt{2 \pi} \sigma} \exp \left(-\frac{\left(y^{(i)}-\theta^{T} x^{(i)}\right)^{2}}{2 \sigma^{2}}\right) \end{aligned} $$To summarize: Under the previous probabilistic assumptions on the data,
least-squares regression corresponds to finding the maximum likelihood estimate of θ. This is thus one set of assumptions under which least-squares regression can be justified as a very natural method that’s just doing maximum likelihood estimation.
(Note however that the probabilistic assumptions are by no means necessary for least-squares to be a perfectly good and rational procedure, and there may—and indeed there are—other natural assumptions that can also be used to justify it.)
We need to minimize $$ \mathcal{J}(\mbf{\theta};\mbf{x},y)= \frac{1}{2} \sum_{i=1}^{n}\mathcal{L}\big(y_{i}, f_{\boldsymbol{\theta}}(\mbf{x}_i)\big)$$ so to find:
$$\mbf{\theta}^{\star} = \arg\min_{\mbf{\theta}} \mathcal{J}(\mbf{\theta};\mbf{x},y) $$In general, if you can find a closed form solution, that is the best you can do.
So if your problem is as simple as inverting a linear system, please **invert a linear system and use pseudo-inverse if you need to!**
In case you cannot derive, we can use numerical, iterative methods
%matplotlib inline
from sklearn import linear_model, datasets
from matplotlib import pyplot as plt
import numpy as np
n_samples = 100
size = 7
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=1,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot()
# Linear Regression
bias = np.ones((X.shape[0], 1))
X = np.hstack((X, bias))
theta = np.linalg.inv(X.T@X)@X.T@y
# Now MeshGrid
x_interp = np.linspace(Xmin, Xmax, 100)
x_interp = x_interp.reshape(-1,1)
x_interp = np.c_[x_interp,np.ones_like(x_interp)]
y_interp = np.dot(theta, x_interp.T)
ax.scatter(x_interp[:,0], y_interp, alpha=0.7, marker='.')
ax.scatter(X[..., 0], y, c='red', marker='o')
ax.set_xlabel('$x$');
ax.set_ylabel('$y$');
%matplotlib inline
from sklearn import linear_model, datasets
from matplotlib import pyplot as plt
import numpy as np
n_samples = 100
size = 7
X, y, coef_gt = datasets.make_regression(
n_samples=n_samples,
n_features=1,
n_informative=1,
noise=20,
coef=True,
random_state=42,
)
fig = plt.figure(figsize=(size, size))
ax = fig.add_subplot()
# Linear Regression
bias = np.ones((X.shape[0], 1))
X = np.hstack((X, bias))
theta = np.linalg.inv(X.T@X)@X.T@y
# Now MeshGrid
x_interp = np.linspace(Xmin, Xmax, 100)
x_interp = x_interp.reshape(-1,1)
x_interp = np.c_[x_interp,np.ones_like(x_interp)]
y_interp = np.dot(theta, x_interp.T)
ax.scatter(x_interp[:,0], y_interp, alpha=0.7, marker='.')
ax.scatter(X[..., 0], y, c='red', marker='o')
ax.set_xlabel('$x$');
ax.set_ylabel('$y$');
fig = plt.figure(figsize=(size-1, size-1))
plt.rcParams['axes.grid'] = False
plt.contourf(xxt, yyt, losses, levels=50, cmap='jet');
plt.colorbar()
plt.scatter(coef_gt,0,color='red',marker='o',s=50)
plt.axis('scaled')
plt.xlabel('$theta_1$')
plt.ylabel('$theta_2$');
## Implementation of Gradient Descent for Logistic Regression
import time
%matplotlib notebook
def get_diff(X, theta, y):
return X@theta - y[..., np.newaxis]
def get_loss(diff):
return 0.5*np.dot(diff.T, diff)
def plot_line(plot3, theta):
x_interp = np.linspace(Xmin, Xmax, 100)
x_interp = x_interp.reshape(-1, 1)
x_interp = np.c_[x_interp, np.ones_like(x_interp)]
y_interp = np.dot(x_interp, theta)
if plot3:
plot3.set_xdata(x_interp[:, 0])
plot3.set_ydata(y_interp)
else:
return x_interp, y_interp
plt.ion()
figure, (axes_1, axes_2) = plt.subplots(2, 2, figsize=(9, 9))
plt.rcParams['axes.grid'] = False
ax0, ax1 = axes_1
ax2, ax3 = axes_2
ax0.contourf(xxt, yyt, losses, levels=50, cmap='jet')
ax0.scatter(coef_gt, 0, color='red', marker='o', s=50)
ax0.set_xlabel('$theta_1$',fontsize=18)
ax0.set_ylabel('$theta_2$',fontsize=18)
ax1.set_ylabel('$loss$',fontsize=18)
ax1.set_xlabel('$iter$',fontsize=18)
ax1.set(xlim=(0, 100), ylim=(0, 1.28e6))
ax2.scatter(X[..., 0], y, c='red', marker='o')
ax2.set_xlabel('$x$',fontsize=18)
ax2.set_ylabel('$y$',fontsize=18)
ax3.set_ylabel('$Grad. Norm.$',fontsize=18)
ax3.set_xlabel('$iter$',fontsize=18)
ax3.set(xlim=(0, 100), ylim=(0, 20000))
theta_curr = np.array([[-100, -100]]).T
losses_track = [get_loss(get_diff(X, theta_curr, y))]
grad_norm_track = [1000]
theta_track = np.array(theta_curr)
lr = 1e-3
loss_tol = 10
plot1, = ax0.plot(*theta_curr, color='violet',
marker='.', markersize=5, linestyle='-')
plot2, = ax1.plot(*losses_track, color='blue',
marker='.', markersize=10, linestyle='--')
xi, yi = plot_line(None, theta_curr)
plot3a,plot3b = ax2.plot(xi, yi, color='blue', marker='.',
markersize=3, linestyle='--')
plot4, = ax3.plot(1000, color='blue',
marker='.', markersize=10, linestyle='--')
while True:
diff = get_diff(X, theta_curr, y)
grad = (diff * X).sum(axis=0, keepdims=True).T
theta_curr = theta_curr - lr*grad
theta_track = np.append(theta_track, theta_curr, axis=1)
diff = get_diff(X, theta_curr, y)
losses_track.append(get_loss(diff))
grad_norm_track.append(np.linalg.norm(grad,2))
if abs(losses_track[-2]-losses_track[-1]) < loss_tol:
break
plot1.set_xdata(theta_track[0, :])
plot1.set_ydata(theta_track[1, :])
plot2.set_xdata(range(len(losses_track)))
plot2.set_ydata(losses_track)
plot4.set_xdata(range(len(grad_norm_track[1:])))
plot4.set_ydata(grad_norm_track[1:])
plot_line(plot3a, theta_curr)
plot_line(plot3b, theta_curr)
figure.canvas.draw()
figure.canvas.flush_events()
time.sleep(0.1)
print(*theta_curr)
plt.show()
[46.09610615] [1.80913084]
where
$$\mathcal{L}\big(y, f_{\boldsymbol{\theta}}(\mbf{x})\big) = \big(f_{\boldsymbol{\theta}}(\mbf{x}) - y \big)^2 $$$$\bmf{\hat{\theta}} = \arg \min_{\bmf{\theta}} \frac{1}{2} \sum_{i=1}^n \big(f_{\boldsymbol{\theta}}(\mbf{x}_i) - y_i \big)^2 $$
The function $\mathcal{J}(\mbf{\theta};\mbf{x},y)$ is a convex quadratic function. The Hessian of $\mathcal{J}(\mbf{\theta};\mbf{x},y)$ at any vector $\theta$ is
the positive definite matrix $\mbf{X}^T\mbf{X}$. Since $\mathcal{J}$ is lower bounded and
grows at infinity, there is a minimum.
plt.rcParams['axes.grid'] = False
fig = plt.figure(figsize=(4, 4))
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(xxt, yyt, losses, cmap='jet')
ax.set_xlabel('$theta_1$')
ax.set_zlabel('$J$')
ax.set_ylabel('$theta_2$');
**Idea: make a little step so that locally after each step the cost is lower than before**
Input: Training set $\{\mbf{x}_i,y_i\}$, learning rate $\gamma$, a small value in $\{0.1,\ldots,\texttt{1e-6}\}$.
Set it to all zeros or random initialization from a distribution. |
Summing up over ${\mbf{x}_i \in \mathbb{R}^d}$ scaled by the difference between the prediction and ground-truth
$$\bmf{\theta} \leftarrow \bmf{\theta} -\gamma \sum_{i=1}^n \big(\underbrace{\bmf{\theta}^T\mbf{x}_i- y_i\big)}_{\text{scalar}}\underbrace{\mbf{x}_i}_{\mathbb{R}^d}$$Problem of GD: What happens if $n \mapsto \infty$.
To make a small step we have to go through ALL the training samples. Optimization could be slow for large $n$.
Idea: make an update for each single training sample selected randomly
$$\bmf{\theta} \leftarrow \bmf{\theta} -\gamma \big(\underbrace{\bmf{\theta}^T\mbf{x}_i- y_i\big)}_{\text{scalar}}\underbrace{\mbf{x}_i}_{\mathbb{R}^d} \qquad \text{where} \qquad i\sim \mathcal{U}(0,n)$$# Implementation of Stochastic Gradient Descent for Logistic Regression
import time
%matplotlib notebook
def get_diff(X, theta, y):
return X@theta - y[..., np.newaxis]
def get_loss(diff):
return 0.5*np.dot(diff.T, diff)
def plot_line(plot3, theta):
x_interp = np.linspace(Xmin, Xmax, 100)
x_interp = x_interp.reshape(-1, 1)
x_interp = np.c_[x_interp, np.ones_like(x_interp)]
y_interp = np.dot(x_interp, theta)
if plot3:
plot3.set_xdata(x_interp[:, 0])
plot3.set_ydata(y_interp)
else:
return x_interp, y_interp
plt.ion()
figure, (axes_1, axes_2) = plt.subplots(2, 2, figsize=(7, 7))
plt.rcParams['axes.grid'] = False
ax0, ax1 = axes_1
ax2, ax3 = axes_2
ax0.contourf(xxt, yyt, losses, levels=50, cmap='jet')
ax0.scatter(coef_gt, 0, color='red', marker='o', s=50)
ax0.set_xlabel('$theta_1$', fontsize=18)
ax0.set_ylabel('$theta_2$', fontsize=18)
ax1.set_ylabel('$loss$', fontsize=18)
ax1.set_xlabel('$iter$', fontsize=18)
ax1.set(xlim=(0, 100), ylim=(0, 1.28e6))
ax2.scatter(X[..., 0], y, c='red', marker='o')
ax2.set_xlabel('$x$', fontsize=18)
ax2.set_ylabel('$y$', fontsize=18)
ax3.set_ylabel('$Grad. Norm.$', fontsize=18)
ax3.set_xlabel('$iter$', fontsize=18)
ax3.set(xlim=(0, 100), ylim=(0, 300))
theta_curr = np.array([[-100, -100]]).T
losses_track = [get_loss(get_diff(X, theta_curr, y))]
grad_norm_track = [1000]
theta_track = np.array(theta_curr)
lr = 1e-1
loss_tol = 10
np.random.seed(42)
plot1, = ax0.plot(*theta_curr, color='violet',
marker='.', markersize=5, linestyle='-')
plot2, = ax1.plot(*losses_track, color='blue',
marker='.', markersize=10, linestyle='--')
xi, yi = plot_line(None, theta_curr)
plot3a, plot3b = ax2.plot(xi, yi, color='blue', marker='.',
markersize=3, linestyle='--')
plot4, = ax3.plot(1000, color='blue',
marker='.', markersize=10, linestyle='--')
while True:
diff = get_diff(X, theta_curr, y)
# STOCHASTIC PART ########################
idx_sampled = np.random.randint(n_samples)
grad = (diff * X)[idx_sampled, :].T.reshape(-1, 1)
##############################
theta_curr = theta_curr - lr*grad
theta_track = np.append(theta_track, theta_curr, axis=1)
diff = get_diff(X, theta_curr, y)
losses_track.append(get_loss(diff))
grad_norm_track.append(np.linalg.norm(grad, 2))
if abs(losses_track[-2]-losses_track[-1]) < loss_tol:
break
plot1.set_xdata(theta_track[0, :])
plot1.set_ydata(theta_track[1, :])
plot2.set_xdata(range(len(losses_track)))
plot2.set_ydata(losses_track)
plot4.set_xdata(range(len(grad_norm_track[1:])))
plot4.set_ydata(grad_norm_track[1:])
plot_line(plot3a, theta_curr)
plot_line(plot3b, theta_curr)
figure.canvas.draw()
figure.canvas.flush_events()
time.sleep(0.1)
print(*theta_curr)
plt.show()
[42.27801342] [-3.01691851]
To summarize: Under the previous probabilistic assumptions on the data,
least-squares regression corresponds to finding the maximum likelihood estimate of θ. This is thus one set of assumptions under which least-squares regression can be justified as a very natural method that’s just doing maximum likelihood estimation.
(Note however that the probabilistic assumptions are by no means necessary for least-squares to be a perfectly good and rational procedure, and there may—and indeed there are—other natural assumptions that can also be used to justify it.)
We assume relations $f \longleftrightarrow y$ is linear We know $D = \{\mbf{x}_i,y_i\}_{i=1}^n$ and we want to find ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d)$
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \theta_0+ \theta_1\cdot x_1 + \theta_2\cdot x_2 + \ldots + \theta_d\cdot x_d$$So ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d) \in \mathbb{R}^{d+1}$.
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \big(\sum_{i=1}^d\theta_i\cdot x_i\big) + \theta_0$$We can augment each feature to have a bias (intercept term) set to $1$ so that $\mbf{x} \doteq [1,\mbf{x}]$.
Doing so $\mbf{x} \in \mathbb{R}^{d+1}$
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \theta_0\cdot \underbrace{x_0}_{\text{always}~1} + \theta_1\cdot x_1 + \theta_2\cdot x_2 + \ldots + \theta_d\cdot x_d$$So ${\boldsymbol{\theta}}\doteq (\theta_0,\ldots,\theta_d) \in \mathbb{R}^{d+1}$.
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^d\theta_i\cdot x_i = \bmf{\theta}^T\mbf{x} $$With $\mbf{x} = [1,x_1,\ldots,x_d]$ and $\mbf{\theta} = [\theta_0,\theta_1,\ldots,\theta_d]$, we have:
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^d\theta_i\cdot x_i = \bmf{\theta}^T\mbf{x} $$We can have another dimensionality $m$ instead of $d$ by using Basis Functions $\bmf{\phi}(\mbf{x})$.
With $\bmf{\phi}(\mbf{x} = [1,\phi(x_1),\ldots,\phi(x_m)]$ and $\mbf{\theta} = [\theta_0,\theta_1,\ldots,\theta_m]$, we have:
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^m\theta_i\cdot x_i = \bmf{\theta}^T\bmf{\phi}(\mbf{x}) $$For Linear Regression:
then we do not have Linear Regression, and the settings impact the model
Let's consider the one dimensional case. We have already seen that we could change the feature $x$ to add the bias term $\mbf{x}\doteq [x,1]$
\begin{align} \mbf{x} &= \begin{bmatrix} x \\ 1 \end{bmatrix} \rightarrow \quad \bmf{\phi}(\mbf{x}) &= \begin{bmatrix} x^2\\ x \\ 1 \end{bmatrix} \end{align}So input dimension is $d=2$ then output dimension after $\bmf{\phi}(\cdot)$ is $m=3$.
In this case we used a second order polynomial to lift up the features $$\bmf{\phi}_m(\mbf{x}) = x^m$$
Important observation:
Important observation:
def solve_lstq(x, y):
return np.linalg.inv(x.T@x)@x.T@y
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red', marker='o', s=30)
plt.scatter(x_valid, y_valid, c='blue', marker='o', s=30)
theta = solve_lstq(x, y)
x_interp = np.linspace(-support_X*1.5, support_X*1.5, 100).reshape(-1, 1)
y_interp = np.dot(theta, x_interp.T)
plt.plot(x_interp[:, 0], y_interp.T, alpha=0.7, marker='.');
err = np.power(y - np.dot(theta, x.T), 2).mean()
Numerically it seems pretty high!
16.42335475440827
Let's consider the one dimensional case. We have already seen that we could change the feature $x$ to add the bias term $\mbf{x}\doteq [x,1]$
\begin{align} \mbf{x} &= \begin{bmatrix} x \\ 1 \end{bmatrix} \rightarrow \quad \bmf{\phi}(\mbf{x}) &= \begin{bmatrix} x^2\\ x \\ 1 \end{bmatrix} \end{align}So input dimension is $d=1$ then output dimension after $\bmf{\phi}(\cdot)$ is $m=2$.
In this case we used a second order polynomial to lift up the features $$\bmf{\phi}_m(\mbf{x}) = x^m$$
Let's consider the one dimensional case. We have already seen that we could change the feature $x$ to add the bias term $\mbf{x}\doteq [x,1]$
\begin{align} \mbf{x} &= \begin{bmatrix} x \\ 1 \end{bmatrix} \rightarrow \quad \bmf{\phi}(\mbf{x}) &= \begin{bmatrix} x^2\\ x \\ 1 \end{bmatrix} \end{align}We can have another dimensionality $m$ instead of $d$ by using Basis Functions $\bmf{\phi}(\mbf{x})$ .
With $\bmf{\phi}(\mbf{x} = [1,\phi(x_1),\ldots,\phi(x_m)]$ and $\mbf{\theta} = [\theta_0,\theta_1,\ldots,\theta_m]$, we have:
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^m\theta_i\cdot \bmf{\phi}_i(x) = \bmf{\theta}^T\bmf{\phi}(\mbf{x}) $$For Linear Regression:
%matplotlib inline
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red',marker='o', s=30)
theta_q = solve_lstq(xq, y)
x_interp = np.linspace(-support_X*1.5, support_X*1.5, 100).reshape(-1, 1)
x_interp_q = np.c_[x_interp, x_interp**2]
y_interp_q = np.dot(theta_q.T, x_interp_q.T)
plt.plot(x_interp_q[:, 0], y_interp_q.T, alpha=0.7, marker='.');
err = np.power(y - np.dot(theta_q.T, xq.T), 2).mean()
Numerically it seems pretty high!
16.04107222204833
We can have another dimensionality $m$ instead of $d$ by using Basis Functions $\bmf{\phi}(\mbf{x})$ .
With $\bmf{\phi}(\mbf{x} = [1,\phi(x_1),\ldots,\phi(x_m)]$ and $\mbf{\theta} = [\theta_0,\theta_1,\ldots,\theta_m]$, we have:
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^m\theta_i\cdot \bmf{\phi}_i(x) = \bmf{\theta}^T\bmf{\phi}(\mbf{x}) $$For Linear Regression:
%matplotlib inline
xc = np.c_[x, x**2, x**3] # make cubic features
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red', marker='o', s=30);
theta_c = solve_lstq(xc,y)
x_interp_c = np.c_[x_interp, x_interp**2, x_interp**3]
y_interp_c = np.dot(theta_c.T, x_interp_c.T)
plt.plot(x_interp_c[:, 0], y_interp_c.T, alpha=0.7, marker='.');
err = np.power(y - np.dot(theta_c.T, xc.T), 2).mean()
Numerically it seems pretty high!
16.036146988841292
We can have another dimensionality $m$ instead of $d$ by using Basis Functions $\bmf{\phi}(\mbf{x})$.
With $\bmf{\phi}(\mbf{x} = [1,\phi(x_1),\ldots,\phi(x_m)]$ and $\mbf{\theta} = [\theta_0,\theta_1,\ldots,\theta_m]$, we have:
$$ f_{\boldsymbol{\theta}}(\mbf{x}) = \sum_{i=0}^m\theta_i\cdot x_i = \bmf{\theta}^T\bmf{\phi}(\mbf{x}) $$For Linear Regression:
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
import numpy as np
errors = []
errors_valid = []
models = []
for m in range(1, 20):
model = Pipeline([('poly', PolynomialFeatures(degree=m, include_bias=False, interaction_only=False)),
('linear', LinearRegression(fit_intercept=False))])
models.append(model)
model = model.fit(x, y)
x_interp = np.linspace(-support_X-offset_valid,
support_X+offset_valid, 100).reshape(-1, 1)
y_interp = model.predict(x_interp)
y_est = model.predict(x)
errors.append(np.power(y - y_est, 2).mean())
errors_valid.append(np.power(y_valid - model.predict(x_valid), 2).mean())
# Draw
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red', marker='o', s=30)
plt.plot(x_interp, y_interp, alpha=0.7, marker='.')
plt.ylim([-10, 10])
fig, axes = plt.subplots(1,2, figsize=(20, 7))
axes[0].bar(range(1, 20), errors)
axes[1].bar(range(1, 20), errors_valid);
axes[1].set_ylim([0,200])
axes[0].set_ylim([0,200])
m_best = np.argmin(errors_valid)
print(f'M best (polynomial degree is) {m_best+1}')
M best (polynomial degree is) 3
# Draw
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red', marker='.')
plt.scatter(x_valid, y_valid, c='blue', marker='.')
y_interp = models[m_best].predict(x_interp)
plt.plot(x_interp, y_interp, alpha=0.7, marker='.');
plt.ylim([-100,100]);
# Draw
plt.figure(figsize=(7, 7))
plt.scatter(x, y, c='red', marker='.')
plt.scatter(x_valid, y_valid, c='blue', marker='.')
y_interp = models[-1].predict(x_interp)
plt.plot(x_interp, y_interp, alpha=0.7, marker='.');
plt.ylim([-100,100]);
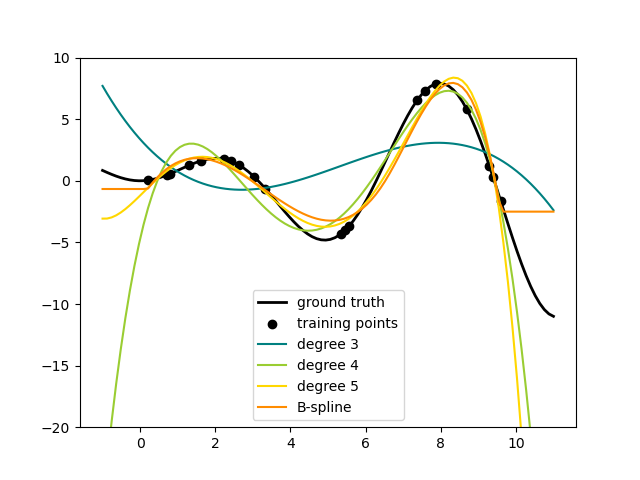
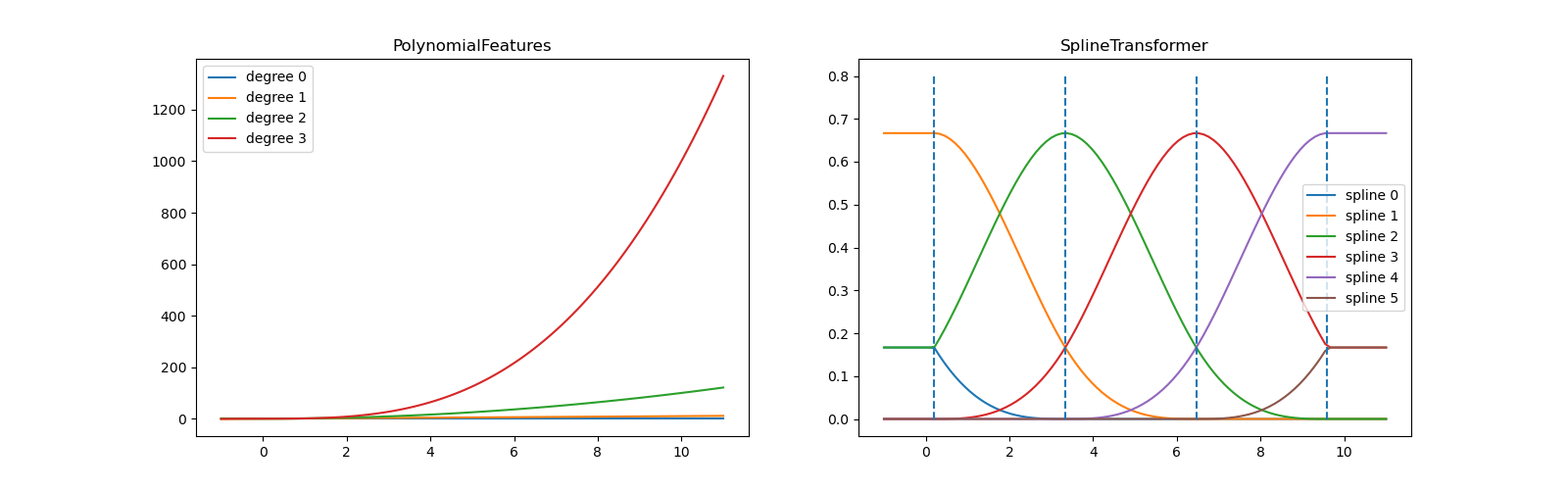
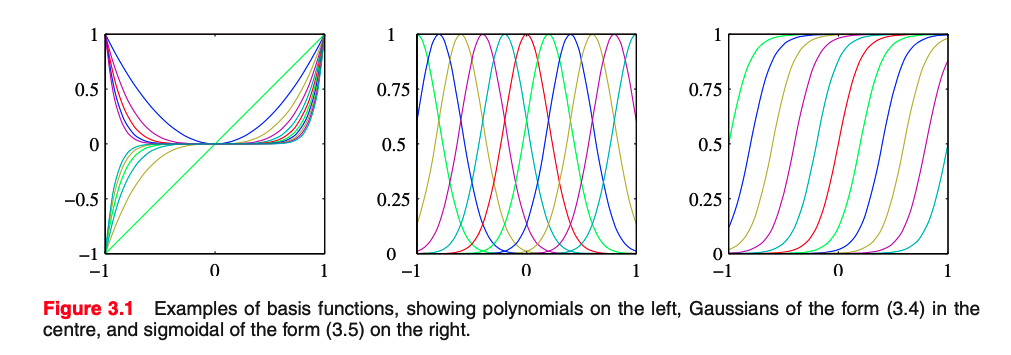
The basis function $\bmf{\phi}_m(\mbf{x}) = x^m$ is global wrt the domain of the feature $x$.
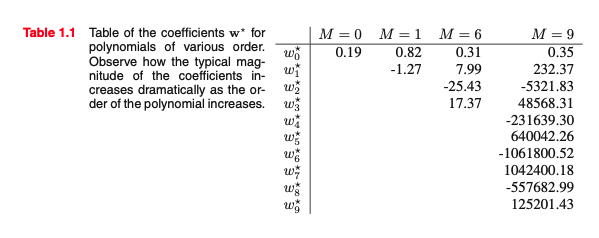
The big limitation of polynomial basis functions is that they are global functions of the input variable, so that changes in one region of input space affect all other regions.
This can be resolved by dividing the input space up into regions and fit a different polynomial in each region, leading to spline functions (that we do not cover).
Advantage: It is simple, and your problem stays convex and well behaved. (i.e. you can still use your original gradient descent code, just with the higher dimensional representation)
Disadvantage: $\phi(x)$ might be Very high dimensional.
X = PolynomialFeatures(interaction_only=True).fit_transform(X)
X_rand = np.array([[3, 7]], dtype=float)
print(f'Input Dimension {X_rand.shape}')
X_rand_poly = PolynomialFeatures(
degree=2, interaction_only=False).fit_transform(X_rand)
print(f'Output Dimension {X_rand_poly.shape}')
print(X_rand[0, :], X_rand_poly[0, :], sep='\n')
Input Dimension (1, 2) Output Dimension (1, 6) [3. 7.] [ 1. 3. 7. 9. 21. 49.]
%matplotlib inline
new_size = [PolynomialFeatures(degree=2, interaction_only=False).fit_transform(
np.random.rand(1, d)).shape[1] for d in range(1, 50)]
plt.figure(figsize=(12,12))
plt.bar(range(1, 50), new_size);

We minimize the cost plus penalty term $$ \mathcal{J}(\mbf{\theta};\mbf{x},y)= \frac{1}{2} \sum_{i=1}^{n}\mathcal{L}\big(y_{i}, f_{\boldsymbol{\theta}}(\mbf{x}_i)\big) + \frac{\lambda}{2} \bmf{\theta}^T \bmf{\theta}= \frac{1}{2} \sum_{i=1}^{n}\mathcal{L}\big(y_{i}, f_{\boldsymbol{\theta}}(\mbf{x}_i)\big) + \frac{\lambda}{2} \vert\vert\bmf{\theta}\vert\vert_2^2 $$ so to find:
$$\mbf{\theta}^{\star} = \arg\min_{\mbf{\theta}} \mathcal{J}_{\text{data}}(\mbf{\theta};\mbf{x},y) +\lambda\mathcal{J}_{\text{reg.}}(\mbf{\theta}) $$Regularization allows complex models to be trained on data sets of limited size without severe over-fitting, essentially by limiting the effective model complexity
Consider the eigendecomposition of the symmetric Positive Semi Definite (PSD) matrix $\mbf{X}\mbf{X}^T$:
$$ X^{T} X= U \Sigma U^{T} =U \underbrace{\left[\begin{array}{ccc} \sigma_{1}^{2} & \ldots & 0 \\ \vdots & \ddots & \vdots \\ 0 & \ldots & \sigma_{d}^{2} \end{array}\right]}_{\operatorname{diag}\left(\sigma_{1}^{2}, \ldots, \sigma_{d}^{2}\right)} U^{T} $$$\sigma_{1}^{2}, \ldots, \sigma_{d}^{2}$ are the eigenvalues and $ UU^{T} = Id$ (since $\Sigma$ is symmetric and square).
If $\mbf{X}$ and $\mbf{X}\mbf{X}^T$ are not full rank then some of $\sigma$ may be zeros.
This implies that if we regularized it, then the pseudo inverse is always invertible and has a unique solution since $\lambda > 0$:
$$ X^{T} X+\lambda I=U\left[\begin{array}{ccc} \sigma_{1}^{2}+\lambda & \ldots & 0 \\ \vdots & \ddots & \vdots \\ 0 & \ldots & \sigma_{d}^{2}+\lambda \end{array}\right] U^{T} $$This implies that if we regularized it then the pseudo inverse is always invertible and has a unique solution since $\lambda > 0$:
$$ \left(X^{T} X+\lambda I\right)^{-1}=U\left[\begin{array}{ccc} \frac{1}{\sigma_{1}^{2}+\lambda} & \cdots & 0 \\ \vdots & \ddots & \vdots \\ 0 & \cdots & \frac{1}{\sigma_{d}^{2}+\lambda} \end{array}\right] U^{T} $$Unlike MLE, now there is a Gaussian prior on the weights; We solve it by doing Maximum A Posteriori (MAP)