


Prof. Iacopo Masi - Computer Science Department, Sapienza University of Rome
Note: Most of the slides are taken from Applied Machine Learning (Cornell CS5785
All the credits go to the authors.
pot and enough deep secret rat thunder black industry answer death material angle crime probable nation debt organization spade acid soup reward free circle west forward board bone substance parcel south scissors move window hanging needle sticky pipe table old river attack design expert$p_{\text{data}(\bx)}$

Picture from [Yang-Song Blog](https://yang-song.net/blog/2021/score/)
$\{\mbf{x}_i\} \sim p_{\text{data}(\bx)}$
Picture from [Yang-Song Blog](https://yang-song.net/blog/2021/score/)
Generative models objective is to learn an approximation of the data density given the data samples (dataset):
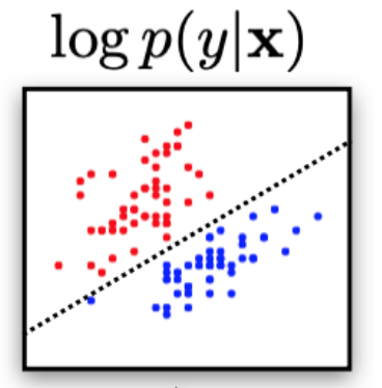
$$ p_{\text{data}}(\bx) \approx p_{\theta}(\bx)$$$$ p_{\text{data}}(\bx,y) \approx p_{\theta}(\bx,y)$$Discriminative models objective is to learn a decision boundary to "separate" data samples to perform classification.
$$p(y == \text{class}~k | \bx)$$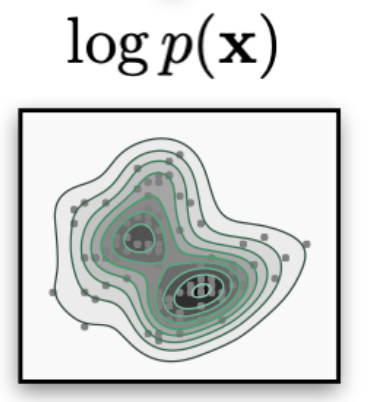
Think "classes" as latent factors in the data--the class label just "reveals" the latent factor. [There could be, there are others]. Think as "slicing the data density" using the latent factor ("coloring" the data density).
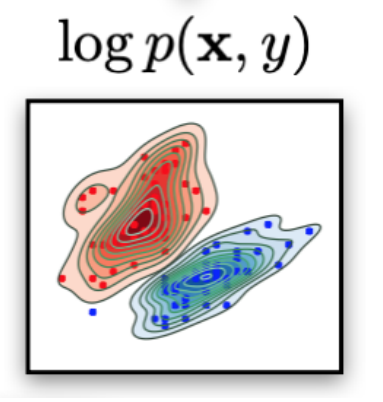
If you need to do just classification (and do not need probabilities):
$$\arg\max_{y^{\prime}} p(y^{\prime}|\bx)$$Then you can:

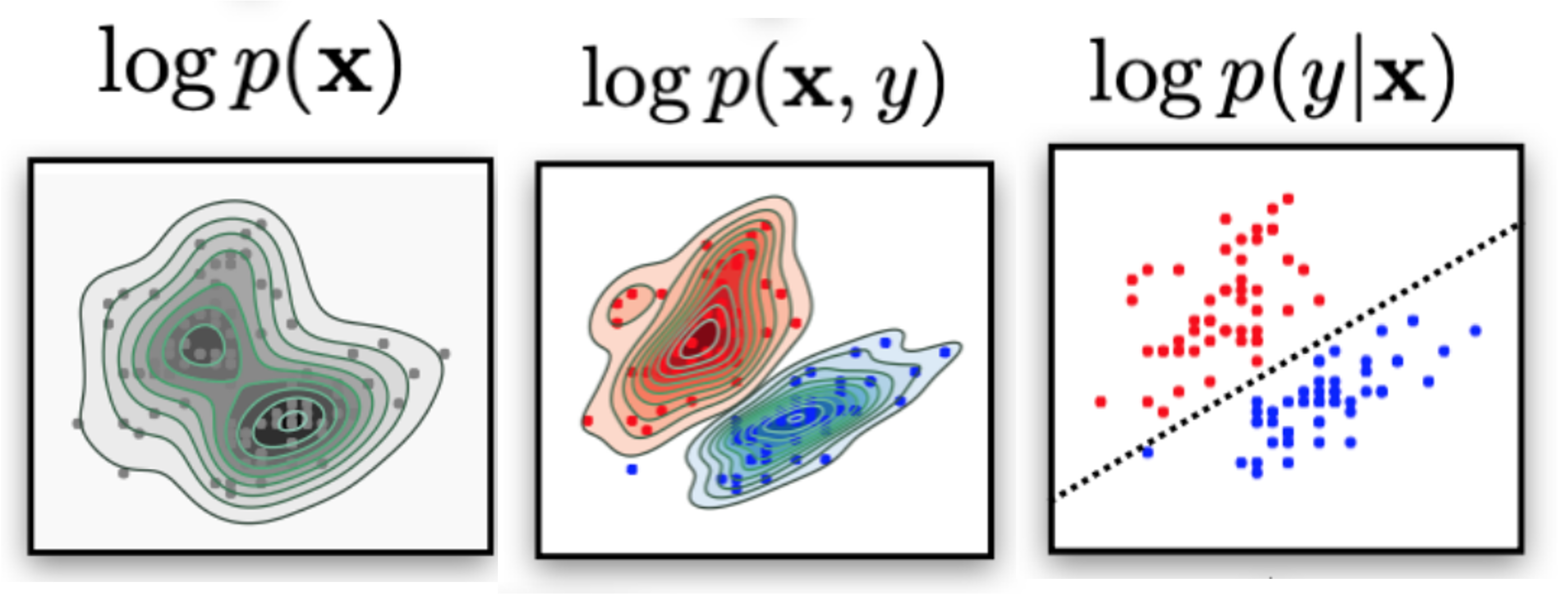
Picture from [YOUR CLASSIFIER IS SECRETLY AN ENERGY BASED MODEL AND YOU SHOULD TREAT IT LIKE ONE](https://openreview.net/pdf?id=Hkxzx0NtDB)
The following slides are taken from Applied Machine Learning (Cornell CS5785
All the credits go to the authors.
Logistic regression is an example of a discriminative classifier. $$ P_\theta(y| x) = \left[ \begin{array}{c} P_\theta(y=0| x) \\ P_\theta(y=1| x) \\ \end{array} \right] $$
Generative classifiers instead define distributions $P_\theta(x|y)$ and $P_\theta(y)$ over $x,y$.
Imagine we have a dataset (e.g., flowers). We can fit a Gaussian to each class:
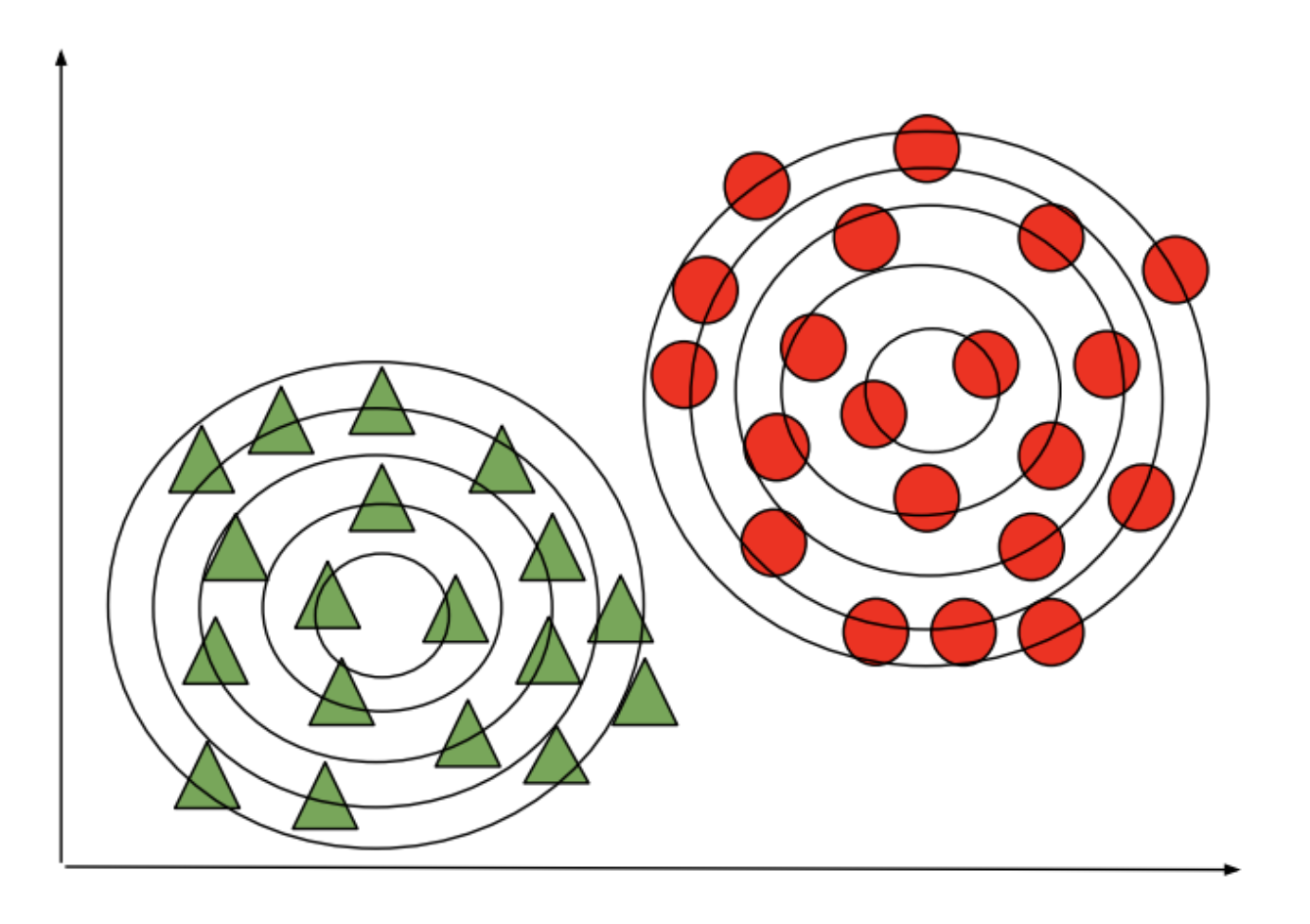
At test time, we predict the class of the Gaussian that is most likely to have generated the data.
Given a new flower $x'$, we would compare the probabilities of three models:
\begin{align*} P_\theta(x'|y=\text{0})P_\theta(y=\text{0}) \\ P_\theta(x'|y=\text{1})P_\theta(y=\text{1}) \\ P_\theta(x'|y=\text{2})P_\theta(y=\text{2}) \end{align*}We output the class that's more likely to have generated $x'$.

Another approach to classification is to use generative models.
In the context of flower classification (e.g., Setosa vs. non-Setosa), we would fit two models on a dataset of flower features $x$ with corresponding species labels $y$:
\begin{align*} P_\theta(x|y=\text{0}) && \text{and} && P_\theta(x|y=\text{1}) \end{align*}We also choose a prior $P(y)$.
Next, we will use GMMs as the basis for a new generative classification algorithm, Gaussian Discriminant Analysis (GDA).
We need to learn the parameters of two sets of distributions:
Next, consider learning $\vec \phi = (\phi_1, \phi_2, \ldots, \phi_K)$.
What are the maximum likelihood $\phi_k$ that are most likely to have generated our data?
Intuitively, the maximum likelihood class probabilities $\phi$ should just be the class proportions that we see in the data. $$ \phi_k = \frac{n_k}{n}$$ for each $k$, where $n_k = |\{i : y^{(i)} = k\}|$ is the number of training targets with class $k$.
Thus, the optimal $\phi_k$ is just the proportion of data points with class $k$ in the training set!
How do we ask the model for predictions? As discussed earlier, we can apply Bayes' rule: $$\arg\max_y P_\theta(y|x) = \arg\max_y P_\theta(x|y)P(y).$$ Thus, we can estimate the probability of $x$ and under each $P_\theta(x|y=k)P(y=k)$ and choose the class that explains the data best.
Let's see how this approach can be used in practice on the Iris dataset.
Let's first start by computing the true parameters on our dataset.
# we can implement these formulas over the Iris dataset
d = 2 # number of features in our toy dataset
K = 3 # number of clases
n = X.shape[0] # size of the dataset
# these are the shapes of the parameters
mus = np.zeros([K,d])
Sigmas = np.zeros([K,d,d])
phis = np.zeros([K])
# we now compute the parameters
for k in range(3):
X_k = X[iris_y == k]
mus[k] = np.mean(X_k, axis=0)
Sigmas[k] = np.cov(X_k.T)
phis[k] = X_k.shape[0] / float(n)
We can look at the learned parameter values and compare them to the data
print('mus:', mus, '\nphis:', phis)
mus: [[5.01 3.43] [5.94 2.77] [6.59 2.97]] phis: [0.33 0.33 0.33]
plt.scatter(X[:, 0], X[:, 1], c=Y, edgecolors='k', cmap=plt.cm.Paired, s=50);
plt.xlabel('Sepal length'); plt.ylabel('Sepal width');
We can compute predictions using Bayes' rule.
# we can implement this in numpy
def gda_predictions(x, mus, Sigmas, phis):
"""This returns class assignments and p(y|x) under the GDA model.
We compute \arg\max_y p(y|x) as \arg\max_y p(x|y)p(y)
"""
# adjust shapes
n, d = x.shape
x = np.reshape(x, (1, n, d, 1))
mus = np.reshape(mus, (K, 1, d, 1))
Sigmas = np.reshape(Sigmas, (K, 1, d, d))
# compute probabilities
py = np.tile(phis.reshape((K,1)), (1,n)).reshape([K,n,1,1])
pxy = (
np.sqrt(np.abs((2*np.pi)**d*np.linalg.det(Sigmas))).reshape((K,1,1,1))
* -.5*np.exp(
np.matmul(np.matmul((x-mus).transpose([0,1,3,2]), np.linalg.inv(Sigmas)), x-mus)
)
)
pyx = pxy * py
return pyx.argmax(axis=0).flatten(), pyx.reshape([K,n])
idx, pyx = gda_predictions(X, mus, Sigmas, phis)
print(idx)
[0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 2 2 2 1 2 1 2 1 2 1 1 1 1 1 1 2 1 1 1 1 1 1 2 1 2 2 2 2 1 1 1 1 1 1 1 2 2 2 1 1 1 1 1 1 1 1 1 1 1 1 2 1 2 2 2 2 1 2 2 2 2 2 2 1 1 2 2 2 2 1 2 1 2 1 2 2 1 1 2 2 2 2 2 1 1 2 2 2 1 2 2 2 1 2 2 2 2 2 2 1]
We visualize the decision boundaries like we did earlier.
Z, pyx = gda_predictions(np.c_[xx.ravel(), yy.ravel()], mus, Sigmas, phis)
logpy = np.log(-1./3*pyx)
Z = Z.reshape(xx.shape)
contours = np.zeros([K, xx.shape[0], xx.shape[1]])
for k in range(K): contours[k] = logpy[k].reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Paired)
for k in range(K): plt.contour(xx, yy, contours[k], levels=np.logspace(0, 1, 1))
plt.scatter(X[:, 0], X[:, 1], c=iris_y, edgecolors='k', cmap=plt.cm.Paired)
plt.xlabel('Sepal length'); plt.ylabel('Sepal width');
Many important generative algorithms are special cases of GDA:
We conclude our lecture on generative classifiers by revisiting the question of how they compare to discriminative classifiers.
When the covariances $\Sigma_k$ in GDA are equal, we have an algorithm called Linear Discriminant Analysis or LDA.
The probability of the data $x$ for each class is a multivariate Gaussian with the same covariance $\Sigma$. $$P_\theta(x|y=k) = \mathcal{N}(x ; \mu_k, \Sigma).$$
The probability over $y$ is Categorical: $P_\theta(y=k) = \phi_k$.
Let's try this algorithm on the Iris flower dataset.
We compute the model parameters similarly to how we did for GDA.
# we can implement these formulas over the Iris dataset
d = 2 # number of features in our toy dataset
K = 3 # number of clases
n = X.shape[0] # size of the dataset
# these are the shapes of the parameters
mus = np.zeros([K,d])
Sigmas = np.zeros([K,d,d])
phis = np.zeros([K])
# we now compute the parameters
for k in range(3):
X_k = X[iris_y == k]
mus[k] = np.mean(X_k, axis=0)
Sigmas[k] = np.cov(X.T) # this is now X.T instead of X_k.T
phis[k] = X_k.shape[0] / float(n)
# print out the means
print(mus)
[[5.01 3.43] [5.94 2.77] [6.59 2.97]]
We can compute predictions using Bayes' rule.
# we can implement this in numpy
def gda_predictions(x, mus, Sigmas, phis):
"""This returns class assignments and p(y|x) under the GDA model.
We compute \arg\max_y p(y|x) as \arg\max_y p(x|y)p(y)
"""
# adjust shapes
n, d = x.shape
x = np.reshape(x, (1, n, d, 1))
mus = np.reshape(mus, (K, 1, d, 1))
Sigmas = np.reshape(Sigmas, (K, 1, d, d))
# compute probabilities
py = np.tile(phis.reshape((K,1)), (1,n)).reshape([K,n,1,1])
pxy = (
np.sqrt(np.abs((2*np.pi)**d*np.linalg.det(Sigmas))).reshape((K,1,1,1))
* -.5*np.exp(
np.matmul(np.matmul((x-mus).transpose([0,1,3,2]), np.linalg.inv(Sigmas)), x-mus)
)
)
pyx = pxy * py
return pyx.argmax(axis=0).flatten(), pyx.reshape([K,n])
idx, pyx = gda_predictions(X, mus, Sigmas, phis)
print(idx)
[0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 2 2 2 1 2 1 2 1 2 1 1 1 1 1 1 2 1 1 1 1 2 1 1 1 2 2 2 2 1 1 1 1 1 1 1 2 2 1 1 1 1 1 1 1 1 1 1 1 1 1 2 1 2 2 2 2 1 2 1 2 2 1 2 1 1 2 2 2 2 1 2 1 2 1 2 2 1 1 2 2 2 2 2 1 1 2 2 2 1 2 2 2 1 2 2 2 1 2 2 1]
We visualize predictions like we did earlier.
Z, pyx = gda_predictions(np.c_[xx.ravel(), yy.ravel()], mus, Sigmas, phis)
logpy = np.log(-1./3*pyx)
Z = Z.reshape(xx.shape)
contours = np.zeros([K, xx.shape[0], xx.shape[1]])
for k in range(K):
contours[k] = logpy[k].reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Paired)
for k in range(K):
plt.contour(xx, yy, contours[k], levels=np.logspace(0, 1, 1))
plt.scatter(X[:, 0], X[:, 1], c=iris_y, edgecolors='k', cmap=plt.cm.Paired)
plt.xlabel('Sepal length'); plt.ylabel('Sepal width');
Linear Discriminant Analysis outputs decision boundaries that are linear, just like Logistic/Softmax Regression.
Softmax or Logistic regression also produce linear boundaries. In fact, both types of algorithms make use of the same model class.
Are the algorithms equivalent?
No! They assume the same model class $\mathcal{M}$, they use a different objective $J$ to select a model in $\mathcal{M}$.
nrow, ncol = 1, 2
fig, axs = plt.subplots(nrow, ncol, figsize=(6*ncol, 2*nrow))
axs[0].pcolormesh(xx, yy, Z, cmap=plt.cm.Paired); axs[0].set_title('LDA')
axs[1].pcolormesh(xx, yy, Z_lr, cmap=plt.cm.Paired); axs[1].set_title('Logistic Regression')
for ax in axs:
ax.scatter(X[:, 0], X[:, 1], c=iris_y, edgecolors='k', cmap=plt.cm.Paired)
ax.set_xlabel('Sepal length'); ax.set_ylabel('Sepal width')
What are the differences between LDA/NB and logistic regression?
Discriminative algorithms are deservingly very popular.
Generative models can also do things that discriminative models can't do.
Given a new $x'$, we would compare the probabilities of both models:
\begin{align*} P_\theta(x'|y=\text{0})P_\theta(y=\text{0}) && \text{vs.} && P_\theta(x'|y=\text{1})P_\theta(y=\text{1}) \end{align*}We output the class that's more likely to have generated $x'$.
We will now do a quick detour to talk about an important application area of machine learning: text classification.
Afterwards, we will see how text classification motivates new classification algorithms.
An interesting instance of a classification problem is classifying text.
What do we do?
A widely-used approach to representing text documents is called "bag of words".
We start by defining a vocabulary $V$ containing all the possible words we are interested in, e.g.: $$ V = \{\text{church}, \text{doctor}, \text{fervently}, \text{purple}, \text{slow}, ...\} $$
A bag of words representation of a document $x$ is a function $\phi(x) \to \{0,1\}^{|V|}$ that outputs a feature vector $$ \phi(x) = \left( \begin{array}{c} 0 \\ 1 \\ 0 \\ \vdots \\ 1 \\ \vdots \\ \end{array} \right) \begin{array}{l} \;\text{church} \\ \;\text{doctor} \\ \;\text{fervently} \\ \vdots \\ \;\text{purple} \\ \vdots \\ \end{array} $$ of dimension $V$. The $j$-th component $\phi(x)_j$ equals $1$ if $x$ contains the $j$-th word in $V$ and $0$ otherwise.
To illustrate the text classification problem, we will use a popular dataset called 20-newsgroups.
Let's load this dataset.
#https://scikit-learn.org/stable/tutorial/text_analytics/working_with_text_data.html
from IPython.display import Markdown, display
import numpy as np; np.set_printoptions(precision=2)
import pandas as pd; pd.options.display.float_format = "{:,.2f}".format
from sklearn.datasets import fetch_20newsgroups
# for this lecture, we will restrict our attention to just 4 different newsgroups:
categories = ['alt.atheism', 'soc.religion.christian', 'comp.graphics', 'sci.med']
# load the dataset
twenty_train = fetch_20newsgroups(subset='train', categories=categories, shuffle=True, random_state=42)
# print some information on it
Markdown(twenty_train.DESCR[:1088])
.. _20newsgroups_dataset:
The 20 newsgroups dataset comprises around 18000 newsgroups posts on 20 topics split in two subsets: one for training (or development) and the other one for testing (or for performance evaluation). The split between the train and test set is based upon a messages posted before and after a specific date.
This module contains two loaders. The first one,
:func:sklearn.datasets.fetch_20newsgroups,
returns a list of the raw texts that can be fed to text feature
extractors such as :class:~sklearn.feature_extraction.text.CountVectorizer
with custom parameters so as to extract feature vectors.
The second one, :func:sklearn.datasets.fetch_20newsgroups_vectorized,
returns ready-to-use features, i.e., it is not necessary to use a feature
extractor.
Data Set Characteristics:
================= ==========
Classes 20
Samples total 18846
Dimensionality 1
Features text
================= ==========# The set of targets in this dataset are the newgroup topics:
twenty_train.target_names
['alt.atheism', 'comp.graphics', 'sci.med', 'soc.religion.christian']
# Let's examine one data point
print(twenty_train.data[3])
From: s0612596@let.rug.nl (M.M. Zwart)
Subject: catholic church poland
Organization: Faculteit der Letteren, Rijksuniversiteit Groningen, NL
Lines: 10
Hello,
I'm writing a paper on the role of the catholic church in Poland after 1989.
Can anyone tell me more about this, or fill me in on recent books/articles(
in english, german or french). Most important for me is the role of the
church concerning the abortion-law, religious education at schools,
birth-control and the relation church-state(government). Thanx,
Masja,
"M.M.Zwart"<s0612596@let.rug.nl>
Let's see an example of Bag-of-Words on 20-newsgroups.
We start by computing these features using the sklearn library.
from sklearn.feature_extraction.text import CountVectorizer
# vectorize the training set
count_vect = CountVectorizer(binary=True)
X_train = count_vect.fit_transform(twenty_train.data)
X_train.shape
(2257, 35788)
The sklearn count vectorizer gives us mappings between words and indices:
print('Word to index `.vocabulary`: ', str(count_vect.vocabulary_)[:94] + ' ...')
feature_names = count_vect.get_feature_names_out()
print('Index to word: ', feature_names[:10])
Word to index `.vocabulary`: {'from': 14887, 'sd345': 29022, 'city': 8696, 'ac': 4017, 'uk': 33256, 'michael': 21661, 'coll ...
Index to word: ['00' '000' '0000' '0000001200' '000005102000' '0001' '000100255pixel'
'00014' '000406' '0007']
We can retrieve the index of $\phi(x)$ associated with each word using these mappings:
# The CountVectorizer class records the index j associated with each word in V
print('Index for the word "church": ', count_vect.vocabulary_.get(u'church'))
print('Index for the word "computer": ', count_vect.vocabulary_.get(u'computer'))
# And we can map the indices back to words
print("Word for the index '8609': ", feature_names[8609])
print("Word for the index '10000': ", feature_names[10000])
Index for the word "church": 8609 Index for the word "computer": 9338 Word for the index '8609': church Word for the index '10000': counseling
Our featurized dataset is in the matrix X_train. We can use the above indices to retrieve the 0-1 value that has been computed for each word:
# We can examine if any of these words are present in our previous datapoint
print(twenty_train.data[3])
# let's see if it contains these two words?
print('---'*20)
print('Value at the index for the word "church": ', X_train[3, count_vect.vocabulary_.get(u'church')])
print('Value at the index for the word "computer": ', X_train[3, count_vect.vocabulary_.get(u'computer')])
print('Value at the index for the word "doctor": ', X_train[3, count_vect.vocabulary_.get(u'doctor')])
print('Value at the index for the word "important": ', X_train[3, count_vect.vocabulary_.get(u'important')])
From: s0612596@let.rug.nl (M.M. Zwart)
Subject: catholic church poland
Organization: Faculteit der Letteren, Rijksuniversiteit Groningen, NL
Lines: 10
Hello,
I'm writing a paper on the role of the catholic church in Poland after 1989.
Can anyone tell me more about this, or fill me in on recent books/articles(
in english, german or french). Most important for me is the role of the
church concerning the abortion-law, religious education at schools,
birth-control and the relation church-state(government). Thanx,
Masja,
"M.M.Zwart"<s0612596@let.rug.nl>
------------------------------------------------------------
Value at the index for the word "church": 1
Value at the index for the word "computer": 0
Value at the index for the word "doctor": 0
Value at the index for the word "important": 1
In practice, we may use some additional modifications of this technique:
Let's now have a look at the performance of classification over bag of words features.
Now that we have a feature representation $\phi(x)$, we can apply the classifier of our choice, such as logistic regression.
from sklearn.linear_model import LogisticRegression
# Create an instance of Softmax and fit the data.
logreg = LogisticRegression(C=1e5, multi_class='multinomial', verbose=True)
logreg.fit(X_train, twenty_train.target)
[Parallel(n_jobs=1)]: Using backend SequentialBackend with 1 concurrent workers. This problem is unconstrained.
RUNNING THE L-BFGS-B CODE
* * *
Machine precision = 2.220D-16
N = 143156 M = 10
At X0 0 variables are exactly at the bounds
At iterate 0 f= 3.12887D+03 |proj g|= 2.12500D+02
* * *
Tit = total number of iterations
Tnf = total number of function evaluations
Tnint = total number of segments explored during Cauchy searches
Skip = number of BFGS updates skipped
Nact = number of active bounds at final generalized Cauchy point
Projg = norm of the final projected gradient
F = final function value
* * *
N Tit Tnf Tnint Skip Nact Projg F
***** 38 43 1 0 0 9.100D-05 1.095D-02
F = 1.0948389171855248E-002
CONVERGENCE: NORM_OF_PROJECTED_GRADIENT_<=_PGTOL
[Parallel(n_jobs=1)]: Done 1 out of 1 | elapsed: 0.6s finished
LogisticRegression(C=100000.0, multi_class='multinomial', verbose=True)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
LogisticRegression(C=100000.0, multi_class='multinomial', verbose=True)
And now we can use this model for predicting on new inputs.
docs_new = ['God is love', 'OpenGL on the GPU is fast']
X_new = count_vect.transform(docs_new)
predicted = logreg.predict(X_new)
for doc, category in zip(docs_new, predicted):
print('%r => %s' % (doc, twenty_train.target_names[category]))
'God is love' => soc.religion.christian 'OpenGL on the GPU is fast' => comp.graphics
In this lecture, we are going to look at generative classifiers and their applications to text classification.
Imagine we have a dataset of, e.g., flowers. We fit a Gaussian to each class:

At test time, we predict the class of the Gaussian that is most likely to have generated the data.
Suppose we want to train a classifier over Iris flowers. In GDA, we define Gaussian probabilities:
\begin{align*} P_\theta(x|y=\text{setosa}) && P_\theta(x|y=\text{non-setosa}) \end{align*}as well as prior probabilities $P_\theta(y=\text{setosa}), P_\theta(y=\text{non-setosa})$.
Each Gaussian $P_\theta(x|y=\text{setosa/non-setosa})$ matches the mean and variance of the class. The $P(y)$ measures the frequency of each class $y$.
Can we apply GDA to text classification?
Next, we are going to look at Naive Bayes, a generative classification algorithm.
We will apply Naive Bayes to the text classification problem.
In binary text classification, we fit two models on a labeled corpus: \begin{align*} P_\theta(x|y=\text{0}) && \text{and} && P_\theta(x|y=\text{1}) \end{align*}
Each model $P_\theta(x | y=k)$ scores $x$ based on how much it looks like class $k$. The documents $x$ are in bag-of-words representation.
How do we choose $P_\theta(x|y=k)$?
A Categorical distribution with parameters $\theta$ is a probability over $K$ discrete outcomes $x \in \{1,2,...,K\}$:
$$ P_\theta(x = j) = \theta_j. $$When $K=2$ this is called the Bernoulli.
A first solution is to assume that $P(x|y=\text{spam})$ is a categorical distribution that assigns a probability to each possible word $x'=[0,1,0,...,0]$: $$ P(x=x'|y=\text{spam}) = P \left( \begin{array}{c} 0 \\ 1 \\ 0 \\ \vdots \\ 0 \end{array} \right. \left. \begin{array}{l} \;\text{church} \\ \;\text{doctor} \\ \;\text{fervently} \\ \vdots \\ \;\text{purple} \end{array} \right) = \theta_{x'} = 0.0012 $$ The $\theta_{x'}$ is the probability of $x'$ under class $\text{spam}$. We need to specify $\theta_{x}$ for all $x$.
How many parameters does a Categorical model $P(x|y=\text{spam})$ have?
For comparison, there are $\approx 10^{82}$ atoms in the universe.
The Naive Bayes assumption is a general technique that can be used with any $d$-dimensional $x$ to construct tractable models $P_\theta(x|y)$ as follows:
$$ P_{\theta} \left( x= \begin{array}{c} 0 \\ 1 \\ 0 \\ \vdots \\ 0 \end{array} \right. \left. \begin{array}{l} \;\text{church} \\ \;\text{doctor} \\ \;\text{fervently} \\ \vdots \\ \;\text{purple} \end{array} \right) = P_\theta(x_1=0) \cdot P_\theta(x_2=1) \cdot ... \cdot P_\theta(x_d=0) $$Each $P_\theta(x_j=0)$ is a Bernoulli.
How many parameters do we need to specify this probability?
$$ P_{\theta} \left( x= \begin{array}{c} 0 \\ 1 \\ 0 \\ \vdots \\ 0 \end{array} \right. \left. \begin{array}{l} \;\text{church} \\ \;\text{doctor} \\ \;\text{fervently} \\ \vdots \\ \;\text{purple} \end{array} \right) = P_\theta(x_1=0) \cdot P_\theta(x_2=1) \cdot ... \cdot P_\theta(x_d=0) $$This typically makes the number of parameters linear instead of exponential in $d$.
To deal with high-dimensional $x$, we choose a simpler model for $P_\theta(x|y)$:
How many parameters does this new model have?
Thus, we only need $Kd$ parameters instead of $K(2^d-1)$!
Naive Bayes assumes that words are uncorrelated, but in reality they are.
As a result, the probabilities estimated by Naive Bayes can be over- under under-confident.
In practice, however, Naive Bayes is a very useful assumption that gives very good classification accuracy!
The Bernoulli Naive Bayes model $P_\theta(x,y)$ is defined for binary data $x \in \{0,1\}^d$ (e.g., bag-of-words documents).
The $\theta$ contains prior parameters $\vec\phi = (\phi_1,...,\phi_K)$ and $K$ sets of per-class parameters $\psi_k = (\psi_{1k},...,\psi_{dk})$.
The probability of the data $x$ for each class equals $$P_\theta(x|y=k) = \prod_{j=1}^d P(x_j \mid y=k),$$ where each $P_\theta(x_j \mid y=k)$ is a $\text{Bernoulli}(\psi_{jk})$.
The probability over $y$ is Categorical: $P_\theta(y=k) = \phi_k$.
We will now turn our attention to learning the parameters of the Naive Bayes model and using them to make predictions.
We need to learn the parameters of two sets of distributions:
Consider learning $\vec \phi = (\phi_1, \phi_2, \ldots, \phi_K)$.
What are the maximum likelihood $\phi_k$ that are most likely to have generated our data?
Intuitively, the class probabilities $\phi$ should just be the class proportions that we see in the data. $$ \phi_k = \frac{n_k}{n}$$ for each $k$, where $n_k = |\{i : y^{(i)} = k\}|$ is the number of training targets with class $k$.
Thus, the optimal $\phi_k$ is just the proportion of data points with class $k$ in the training set!
Next we want to learn the parameter $\psi_{jk}$ (i.e., probability that $x_j=1$ given class $k$).
What is the optimal $\psi_{jk}$ in this case?
Each $\psi_{jk}$ is simply the proportion of documents in class $k$ that contain the word $j$. \begin{align*} \psi_{jk} = \frac{n_{jk}}{n_k}. \end{align*} where $|\{i : x^{(i)}_j = 1 \text{ and } y^{(i)} = k\}|$ is the number of $x^{(i)}$ with label $k$ and a positive occurrence of word $j$.
How do we ask the model for predictions? As discussed earlier, we can apply Bayes' rule: $$\arg\max_y P_\theta(y|x) = \arg\max_y P_\theta(x|y)P(y).$$ Thus, we can estimate the probability of $x$ and under each $P_\theta(x|y=k)P(y=k)$ and choose the class that explains the data best.
To illustrate the text classification problem, we will use a popular dataset called 20-newsgroups.
Let's load this dataset.
#https://scikit-learn.org/stable/tutorial/text_analytics/working_with_text_data.html
from sklearn.datasets import fetch_20newsgroups
# for this lecture, we will restrict our attention to just 4 different newsgroups:
categories = ['alt.atheism', 'soc.religion.christian', 'comp.graphics', 'sci.med']
twenty_train = fetch_20newsgroups(subset='train', categories=categories, shuffle=True, random_state=42)
Markdown(twenty_train.DESCR[:1088]);
Let's see an example of Naive Bayes on 20-newsgroups.
We start by computing these features using the sklearn library.
from sklearn.feature_extraction.text import CountVectorizer
# vectorize the training set
count_vect = CountVectorizer(binary=True, max_features=1000)
y_train = twenty_train.target
X_train = count_vect.fit_transform(twenty_train.data).toarray()
feature_names = count_vect.get_feature_names_out()
X_train.shape
(2257, 1000)
Let's compute the maximum likelihood model parameters on our dataset. This is done in closed-form by just estimating statistics on the data, so we don't need to do gradient descent.
n = X_train.shape[0] # size of the dataset
d = X_train.shape[1] # number of features in our dataset
K = 4 # number of clases
# these are the shapes of the parameters
psis = np.zeros([K,d])
phis = np.zeros([K])
# we now compute the parameters
for k in range(K):
X_k = X_train[y_train == k]
psis[k] = np.mean(X_k, axis=0)
phis[k] = X_k.shape[0] / float(n)
# print out the class proportions
print(phis)
[0.21 0.26 0.26 0.27]
The learned parameters contain information about the most frequent words:
top_words = []
for category in range(4):
top_words.append(feature_names[np.argsort(psis[category])[-200:]])
for category in range(4):
words_from_other_categories = np.concatenate(top_words[:category] + top_words[category+1:])
unique_words = [word for word in top_words[category] if word not in words_from_other_categories]
print(f'\n# top 10 ~unique words occurring in {twenty_train.target_names[category]}:')
print(unique_words[-10:])
# top 10 ~unique words occurring in alt.atheism: ['someone', 'again', 'allan', 'political', 'schneider', 'atheism', 'caltech', 'cco', 'keith', 'atheists'] # top 10 ~unique words occurring in comp.graphics: ['files', 'version', 'file', 'mail', 'image', 'keywords', 'program', 'looking', 'please', 'graphics'] # top 10 ~unique words occurring in sci.med: ['soon', 'univ', 'pittsburgh', 'disease', 'geb', 'banks', 'medical', 'years', 'gordon', 'pitt'] # top 10 ~unique words occurring in soc.religion.christian: ['faith', 'athos', 'church', 'bible', 'christ', 'jesus', 'apr', 'christians', '1993', 'rutgers']
We can compute predictions using Bayes' rule.
def nb_predictions(x, psis, phis):
"""This returns class assignments and scores under the NB model.
We compute \arg\max_y p(y|x) as \arg\max_y p(x|y)p(y)
"""
# adjust shapes
n, d = x.shape
x = np.reshape(x, (1, n, d))
psis = np.reshape(psis, (K, 1, d))
psis = psis.clip(1e-14, 1-1e-14) # clip probabilities to avoid log(0)
# compute log-probabilities
logpy = np.log(phis).reshape([K,1])
logpxy = x * np.log(psis) + (1-x) * np.log(1-psis)
logpyx = logpxy.sum(axis=2) + logpy
return logpyx.argmax(axis=0).flatten(), logpyx.reshape([K,n])
idx_train, logpyx = nb_predictions(X_train, psis, phis)
print(idx_train[:10])
[1 1 3 0 3 3 3 2 2 2]
We can measure the accuracy:
(idx_train==y_train).mean()
0.8692955250332299
We can also make up new queries and ask the model to label them:
docs_new = ['OpenGL on the GPU is fast']
X_new = count_vect.transform(docs_new).toarray()
predicted, logpyx_new = nb_predictions(X_new, psis, phis)
for doc, category in zip(docs_new, predicted):
print('%r => %s' % (doc, twenty_train.target_names[category]))
'OpenGL on the GPU is fast' => comp.graphics
We conclude by deriving the formula for Naive Bayes from scratch.
We can learn a generative model $P_\theta(x, y)$ by maximizing the maximum likelihood:
$$ \max_\theta \sum_{i=1}^n \log P_\theta({x}^{(i)}, y^{(i)}). $$This seeks to find parameters $\theta$ such that the model assigns high probability to the training data.
Let's use maximum likelihood to fit the Bernoulli Naive Bayes model. Note that model parameters $\theta$ are the union of the parameters of each sub-model: $$\theta = (\phi_1, \phi_2,\ldots, \phi_K, \psi_{11}, \psi_{21}, \ldots, \psi_{dK}).$$
Given a dataset $\mathcal{D} = \{(x^{(i)}, y^{(i)})\mid i=1,2,\ldots,n\}$, we want to optimize the log-likelihood $\ell(\theta) = \log L(\theta)$: \begin{align*} \ell(\theta) & = \sum_{i=1}^n \sum_{j=1}^d \log P_\theta(x^{(i)}_j | y^{(i)}) + \sum_{i=1}^n \log P_\theta(y^{(i)}) \\ & = \sum_{k=1}^K \sum_{j=1}^d \underbrace{\sum_{i :y^{(i)} =k} \log P(x^{(i)}_j | y^{(i)} ; \psi_{jk})}_\text{all the terms that involve $\psi_{jk}$} + \underbrace{\sum_{i=1}^n \log P(y^{(i)} ; \vec \phi)}_\text{all the terms that involve $\vec \phi$}. \end{align*}
In equality #2, we use Naive Bayes: $P_\theta(x,y)=P_\theta(y) \prod_{i=1}^d P(x_j|y)$; in the third one, we change the order of summation.
Each $\psi_{jk}$ for $k=1,2,\ldots,K$ is found in only the following terms: $$ \max_{\psi_{jk}} \ell(\theta) = \max_{\psi_{jk}} \sum_{i :y^{(i)} =k} \log P(x^{(i)}_j | y^{(i)} ; \psi_{jk}). $$ Thus, optimization over $\psi_{jk}$ can be carried out independently of all the other parameters by just looking at these terms.
Similarly, optimizing for $\vec \phi = (\phi_1, \phi_2, \ldots, \phi_K)$ only involves a few terms: $$ \max_{\vec \phi} \sum_{i=1}^n \log P_\theta(x^{(i)}, y^{(i)} ; \theta) = \max_{\vec\phi} \sum_{i=1}^n \log P_\theta(y^{(i)} ; \vec \phi). $$
Let's first consider the optimization over $\vec \phi = (\phi_1, \phi_2, \ldots, \phi_K)$. $$ \max_{\vec \phi} \sum_{i=1}^n \log P_\theta(y=y^{(i)} ; \vec \phi). $$
What is the maximum likelihood $\phi$ in this case?
Intuitively, the maximum likelihood class probabilities $\phi$ should just be the class proportions that we see in the data.
Let's calculate this formally. Our objective $J(\vec \phi)$ equals \begin{align*} J(\vec\phi) & = \sum_{i=1}^n \log P_\theta(y^{(i)} ; \vec \phi) = \sum_{i=1}^n \log \left( \frac{\phi_{y^{(i)}}}{\sum_{k=1}^K \phi_k}\right) \\ & = \sum_{i=1}^n \log \phi_{y^{(i)}} - n \cdot \log \sum_{k=1}^K \phi_k \\ & = \sum_{k=1}^K \sum_{i : y^{(i)} = k} \log \phi_k - n \cdot \log \sum_{k=1}^K \phi_k \end{align*}
Taking the derivative and setting it to zero, we obtain $$ \frac{\phi_k}{\sum_l \phi_l} = \frac{n_k}{n}$$ for each $k$, where $n_k = |\{i : y^{(i)} = k\}|$ is the number of training targets with class $k$.
Thus, the optimal $\phi_k$ is just the proportion of data points with class $k$ in the training set!
Next, let's look at the maximum likelihood term $$ \arg\max_{\psi_{jk}} \sum_{i :y^{(i)} =k} \log P(x^{(i)}_j | y^{(i)} ; \psi_{jk}). $$ over the word parameters $\psi_{jk}$.
What is the maximum likelihood $\psi_{jk}$ in this case?
Each $\psi_{jk}$ is simply the proportion of documents in class $k$ that contain the word $j$.
We can maximize the likelihood exactly like we did for $\phi$ to obtain closed-form solutions: \begin{align*} \psi_{jk} = \frac{n_{jk}}{n_k}. \end{align*} where $|\{i : x^{(i)}_j = 1 \text{ and } y^{(i)} = k\}|$ is the number of $x^{(i)}$ with label $k$ and a positive occurrence of word $j$.
What are the pros and cons of generative and discriminative methods?
Next class: more generative models, beyond the Naive Bayes limitations, fitting mixtures to data