

Like PCA, we can define a loss or cost function that can better formalize the K-means algorithm.
$$ \mathcal{L}(\mbf{\mu},y;\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \mbf{\mu}_{y_i} \right|\right|_2^2 $$Given the scalar function:
$$ \mathcal{L}(\mbf{\mu};\mbf{D}) = \sum_{i=1}^N \left( {x}_i - {\mu} \right)^2 $$Write a proof sketch that ${\mu}$ as the average of $\mbf{D}\doteq\{{x}_i\}_{i=1}^N$ is a critical point of the function. (It is also a global minimum)
Given the function vector to scalar function, where the assignments $\bar{y}$ are given somehow (they cannot "move"):
$$ \mathcal{L}(\mbf{\mu},\bar{y};\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \boldsymbol{\mu}_{\bar{y}_i} \right|\right|_2^2 $$Write a proof sketch that $\boldsymbol{\mu}$ as the average of the vectors $\mbf{D}\doteq\{\mbf{x}_i\}_{i=1}^N$ is a critical point of the function. (It is also a global minimum)
Hint: set the derivative/gradient of mu to zero and solve; for 2b) pay attention to how the L2 norm is defined
Objective and Motivation: The goal of unsupervised learning is to find hidden patterns in unlabeled data.
\begin{equation} \underbrace{\{\mathbf{x}_i\}_{i=1}^N}_{\text{known}} \sim \underbrace{\mathcal{D}}_{\text{unknown}} \end{equation}Unlike in supervised learning, any data point is not paired with a label.
As you can see the unsupervised learning problem is ill-posed (which hidden patterns?) and in principle more difficult than supervised learning.
Unsupervised learning can be thought of as "finding structure" in the data.
dist = np.array([1.18518298, 1.30917493, 1.10973212, 2.24523519, 1.01625606])
to pmf = np.array([0.17262675 0.19068668 0.16163702 0.32702769 0.14802185]) simply with
pmf = dist/dist.sum()
fig, axs = plt.subplots(1, 2)
fig.set_figheight(3)
fig.set_figwidth(15)
# PDF
axs[0].stem(pmf, linefmt='b-', markerfmt='bo', basefmt='--')
axs[0].set_title('PMF')
axs[0].set_xlabel('Index of distance')
axs[0].set_ylabel('Probability')
axs[0].set_aspect('auto')
# CUMSUM
axs[1].plot(pmf.cumsum(), 'o--')
axs[1].set_title('CDF')
axs[1].set_xlabel('Index of distance')
axs[1].set_ylabel('Cumulative Probability')
axs[1].set_aspect('auto')
plt.show()
# Univariate Gaussian (1-D Gaussian)
mu, sigma = 0, 1
x = np.arange(-3, 3, 0.01)
p = 1 / np.sqrt(2 * np.pi * sigma**2) * np.exp(-(x - mu)**2 / (2 * sigma**2))
_ = plt.plot(x, p);
_ = plt.title('Gaussian PDF')
from scipy.stats import multivariate_normal
mu = [0, 0]
Sigma = [[2, 0.0],
[0.0, 1]]
F = multivariate_normal(mu, Sigma )
from scipy.stats import multivariate_normal
X, Y = np.mgrid[-2:2:0.01, -2:2:0.01]
pos = np.dstack((X, Y))
Z = F.pdf(pos)
# plot using subplots
fig = plt.figure(figsize=(20,20))
ax1 = fig.add_subplot(1, 2, 1, projection='3d')
ax1.plot_surface(X, Y, Z, rstride=3, cstride=3, linewidth=1, antialiased=True,
cmap='jet')
ax1.view_init(55, -70)
# ax1.set_xticks([])
# ax1.set_yticks([])
# ax1.set_zticks([])
ax1.set_xlabel(r'$x_1$')
ax1.set_ylabel(r'$x_2$')
ax2 = fig.add_subplot(1, 2, 2, projection='3d')
ax2.contourf(X, Y, Z, zdir='z', offset=0, cmap='jet')
ax2.view_init(90, 270)
ax2.grid(False)
ax2.set_xticks([])
ax2.set_yticks([])
ax2.set_zticks([])
ax2.set_xlabel(r'$x_1$')
ax2.set_ylabel(r'$x_2$')
plt.show()
from scipy.stats import multivariate_normal
step = 0.01
X, Y = np.mgrid[-2:2:step, -2:2:step]
pos = np.dstack((X, Y))
##############################
### Gaussian PDF definition
Sigma = [[.001, 0.0],
[0.0, .001]]
mu = [0, 0]
F = multivariate_normal(mu, Sigma )
##############################
Z = F.pdf(pos)
# plot using subplots
fig = plt.figure(figsize=(20,20))
ax1 = fig.add_subplot(1, 2, 1, projection='3d')
ax1.plot_surface(X, Y, Z, rstride=3, cstride=3, linewidth=1, antialiased=True,
cmap='jet')
ax1.view_init(55, -70)
# ax1.set_xticks([])
# ax1.set_yticks([])
# ax1.set_zticks([])
ax1.set_xlabel(r'$x_1$')
ax1.set_ylabel(r'$x_2$')
ax1.set_xlim3d(-2,2)
ax1.set_ylim3d(-2,2)
#ax1.set_zlim3d(0,0.15)
ax2 = fig.add_subplot(1, 2, 2, projection='3d')
ax2.contourf(X, Y, Z, zdir='z', offset=0, cmap='jet')
ax2.view_init(90, 270)
ax2.grid(False)
ax2.set_xticks([])
ax2.set_yticks([])
ax2.set_zticks([])
ax2.set_xlabel(r'$X_1$')
ax2.set_ylabel(r'$X_2$')
ax2.set_xlim(-2,2)
ax2.set_ylim(-2,2)
plt.show()
The Mahalanobis distance of a point $\mbf{x}$ from a multivariate Gaussian $\mathcal{N}(\mbf{\mu},\mbf{\Sigma})$
$$ D_M(\vec{x}) = \sqrt{(\mbf{x} - \mbf{\mu})^\mathsf{T} \mathbf{\Sigma}^{-1} (\mbf{x} - \mbf{\mu})} $$This distance is zero for $\mbf{x}$ at $\mbf{\mu}$ and grows as $\mbf{x}$ moves away from the mean along each principal component axis.
If each of these axes is re-scaled to have unit variance, then the Mahalanobis distance corresponds to the standard Euclidean distance in the transformed space. The Mahalanobis distance is thus unitless, scale-invariant, and takes into account the correlations of the data set.
Which is also equal to: $$\underbrace{p(X,Y)}_{\text{joint}} = \underbrace{p(Y)}_{\text{marginal}} \underbrace{p(X|Y)}_{\text{cond.}}$$
You can also write the denominator more explicitly by expanding the marginal distribution (discrete case).
$$ p(Y|X) = \frac{p(Y)p(X|Y)}{p(X)} = \frac{p(Y)p(X|Y)}{\sum_{Y^{\prime}}p(Y^{\prime})p(X|Y^{\prime})} $$The joint probability simply becomes the product fo the marginals.
$$ p(X,Y) = p(X)p(Y|X) = p(X)p(Y) \text{ because } p(Y) = p(Y|X)$$when they $X \perp Y $
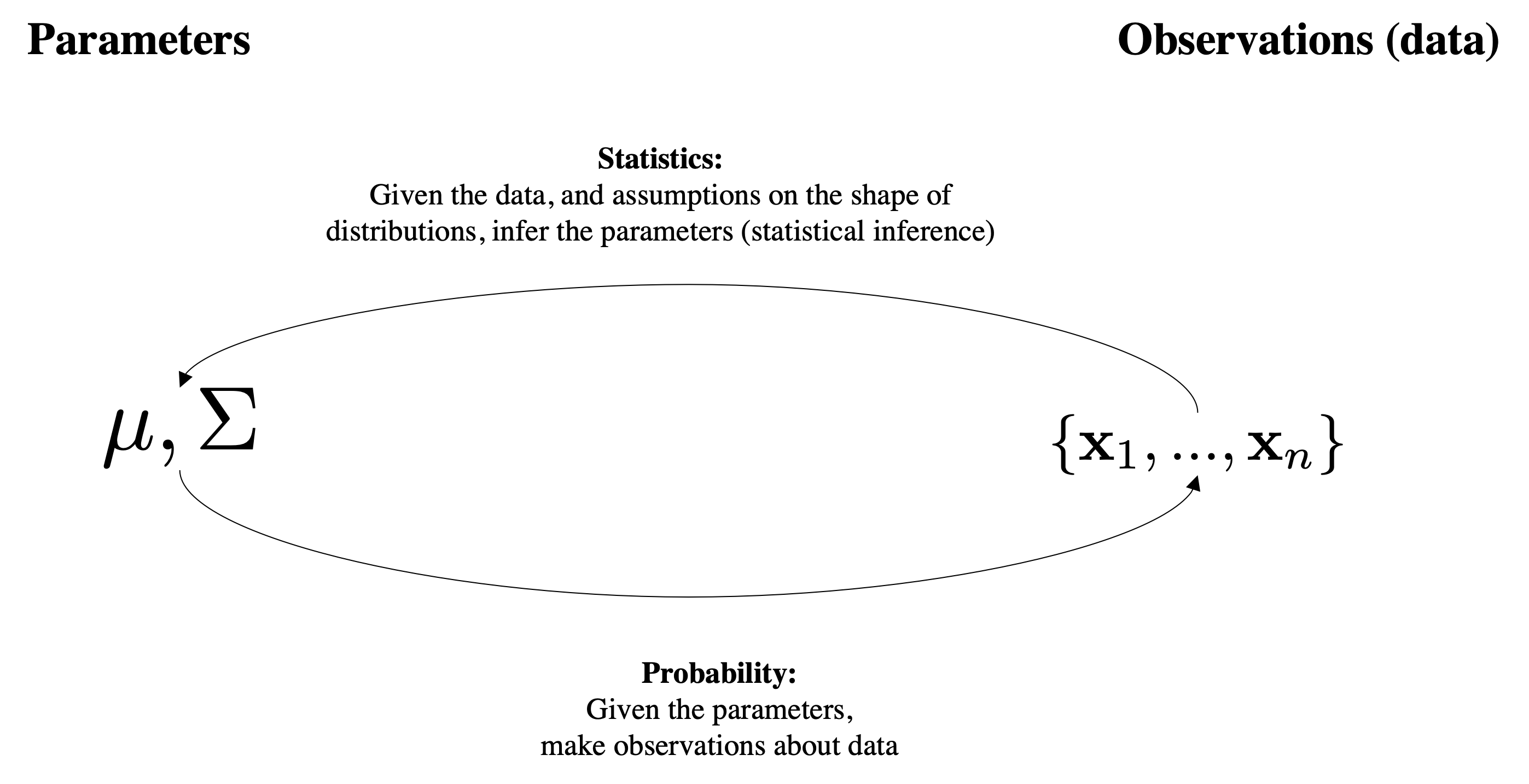
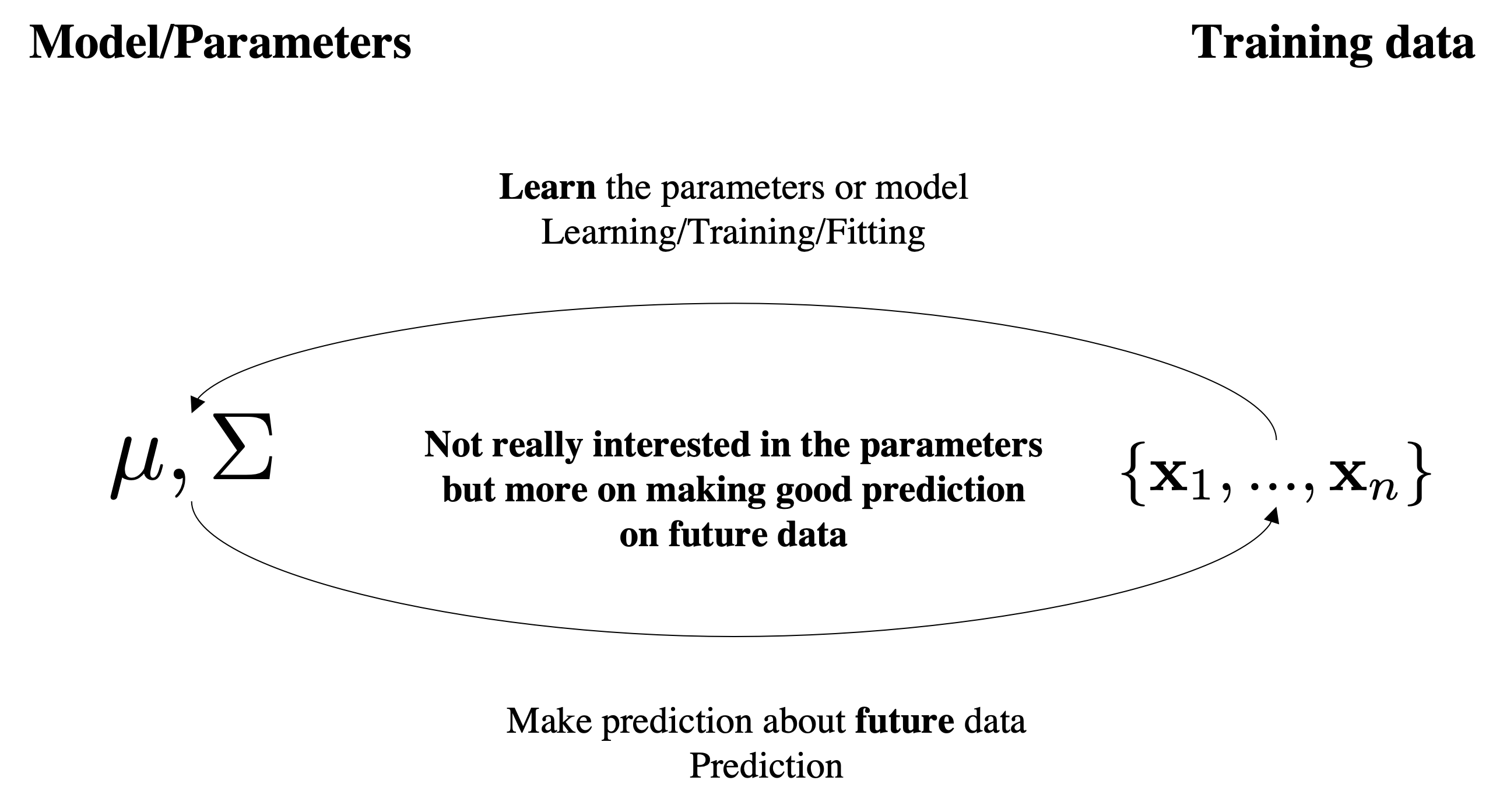
Assumptions
This has a Bayesian interpretation which can be helpful to think about. Suppose that we have a model with parameters $\boldsymbol{\theta}\doteq\mu,\Sigma$ and a collection of data examples $X=\{\mbf{x}_1,\ldots,\mbf{x}_N \}$.
If we want to find the most likely value for the parameters of our model, given the data, that means we want to find
$$\mathop{\mathrm{argmax}}_ {\boldsymbol{\theta}} P(\boldsymbol{\theta}\mid X).$$By Bayes' rule, this is the same thing as
$$ \mathop{\mathrm{argmax}}_ {\boldsymbol{\theta}} P(\boldsymbol{\theta}\mid X) = \mathop{\mathrm{argmax}}_ {\boldsymbol{\theta}} \frac{P(X \mid \boldsymbol{\theta})P(\boldsymbol{\theta})}{P(X)}. $$Thus we see that our application of Bayes' rule shows that our best choice of $\boldsymbol{\theta}$ is the maximum likelihood estimate for $\boldsymbol{\theta}$:
$$ \hat{\boldsymbol{\theta}} = \mathop{\mathrm{argmax}} _ {\boldsymbol{\theta}} P(X \mid \boldsymbol{\theta}) = \mathop{\mathrm{argmax}} _ {\boldsymbol{\theta}} P(\boldsymbol{\theta}\mid X ) $$This is the reason why likelihood and pdf are the same but we just change the way we interpret the variables inside!
As a matter of common terminology, the probability of the data given the parameters $p(X\mid\boldsymbol{\theta})$ is referred to as the likelihood.
This holds also for the likelihood:
$$ L({\color{green}{\mbf{\mu}, \mbf{\Sigma}}};\mbf{x}) = \prod_{i=1}^N \mathcal{N}(\bx_i; {\color{green}{\mbf{\mu}, \mbf{\Sigma}}}) $$We can find maximum points by taking the derivative and setting it to zero: $$ \frac{\partial L({\color{green}{\mbf{\mu}, \mbf{\Sigma}}};\bx)}{\partial {\color{green}{\mbf{\mu}}}} = \mbf{0} $$
We can find maximum points by taking the derivative and setting it to zero: $$ \frac{\partial L({\color{green}{\mbf{\mu}, \mbf{\Sigma}}};\bx)}{\partial {\color{green}{\mbf{\Sigma}}}} = \mbf{0} $$
We use log because:
log and exp and $\prod$ will change to $\sum$ because of log propertiesargmax, not maxor
$$ {\color{green}{\mbf{\theta}^*}} = \arg\max_{\color{green}{\mbf{\theta}}} \left[ \sum_{i=1}^N \ln \big( L({\color{green}{\mbf{\mu}, \mbf{\Sigma}}};\bx_i) \big) \right] $$This holds also for the likelihood:
$$ L({\color{green}{\mbf{\mu}, \mbf{\Sigma}}};\mbf{X}) = \prod_{i=1}^N \mathcal{N}(\bx_i; {\color{green}{\mbf{\mu}, \mbf{\Sigma}}}) $$Using log: $$ \arg\max \sum_{i=1}^N \ln \big( \mathcal{N}(\bx_i; {\color{green}{\mbf{\mu}, \mbf{\Sigma}}}) \big) $$
And replacing Gaussian equation: $$ \arg\max \sum_{i=1}^N \ln \left( \frac{1}{(2 \pi)^{D / 2} |{\color{green}{\mbf{\Sigma}}}|^{1 / 2}} \exp \left\{ -\frac{1}{2}(\bx_i - {\color{green}{\mbf{\mu}}})^{\mathrm{T}} {\color{green}{\mbf{\Sigma}^{-1}}} (\bx_i - {\color{green}{\mbf{\mu}}}) \right\} \right) $$
from scipy.stats import norm
np.random.seed(1)
########################################
mu, sigma, N = 0, 1, 10 # mean and standard deviation of the generative process
# when we work in practice we have only points we do not know the generative pocess.
########################################
points = np.random.normal(mu, sigma, N)
X_plot = np.linspace(-5, 5, 1000)
true_dens = norm(mu, sigma).pdf(X_plot)
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="Input distribution (unknown)")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
ax.legend()
<matplotlib.legend.Legend at 0x7f7881b48b20>
def estimate_gaussian_mle(points, X_plot, plot=True):
# Now, we estimate with MLE in close form
mu_mle = points.mean()
std_mle = np.std(points, ddof=0)
if plot:
MLE_dens = norm(mu_mle, std_mle).pdf(X_plot)
_ = ax.fill(X_plot, MLE_dens, fc="red",
alpha=0.2, label="estimated")
return mu_mle, std_mle
mu_mle, std_mle = estimate_gaussian_mle(points, X_plot)
print(f'Estimated ({mu_mle}, {std_mle}) vs Ground-truth ({mu}, {sigma})')
Estimated (-0.09714089080609985, 1.190898552063902) vs Ground-truth (0, 1)
#help(np.std) #toggle to get help on the standard deviation function in numpy
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="input distribution")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
mu_mle, std_mle = estimate_gaussian_mle(points, X_plot)
print(f'Estimated ({mu_mle}, {std_mle}) vs Ground-truth ({mu}, {sigma})')
Estimated (-0.09714089080609985, 1.190898552063902) vs Ground-truth (0, 1)
from scipy.stats import norm
np.random.seed(1)
########################################
mu, sigma, N = 0, 1, 100 # mean and standard deviation of the generative process
# when we work in practice, we have only points and we do not know the generative process.
########################################
# when we work in practice, we have only points and we do not know the generative process.
points = np.random.normal(mu, sigma, N)
X_plot = np.linspace(-5, 5, 1000)
true_dens = norm(mu, sigma).pdf(X_plot)
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="input distribution")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="input distribution")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
mu_mle, std_mle = estimate_gaussian_mle(points, X_plot)
print(f'Estimated ({mu_mle}, {std_mle}) vs Ground-truth ({mu}, {sigma})')
Estimated (0.060582852075698704, 0.885156213831585) vs Ground-truth (0, 1)
from scipy.stats import norm
np.random.seed(1)
########################################
mu, sigma, N = 0, 1, 1000 # mean and standard deviation of the generative process
# when we work in practice, we have only points and we do not know the generative process.
########################################
# when we work in practice, we have only points and we do not know the generative process.
points = np.random.normal(mu, sigma, N)
X_plot = np.linspace(-5, 5, 1000)
true_dens = norm(mu, sigma).pdf(X_plot)
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="input distribution")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
fig, ax = plt.subplots(figsize=(5, 5))
_ = ax.fill(X_plot, true_dens, fc="black",
alpha=0.2, label="input distribution")
_ = ax.plot(points, -0.005 - 0.01 * np.random.random(points.shape[0]), "+k")
mu_mle, std_mle = estimate_gaussian_mle(points, X_plot)
print(f'Estimated ({mu_mle}, {std_mle}) vs Ground-truth ({mu}, {sigma})')
Estimated (0.03881247615960185, 0.9810041339322116) vs Ground-truth (0, 1)
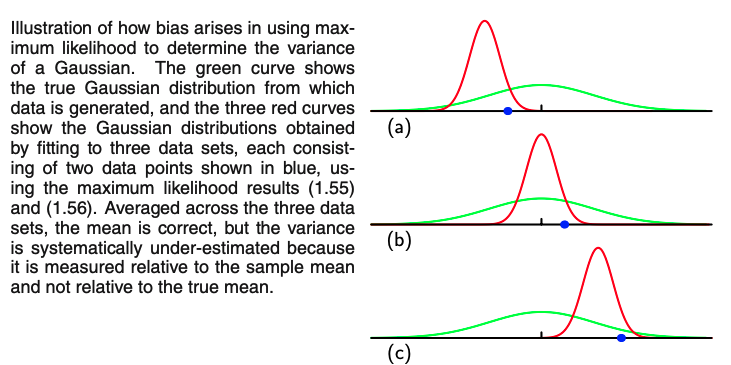
Like PCA, we can define a loss or cost function that can better formalize the K-means algorithm.
$$ \mathcal{L}(\mbf{\mu},y;\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \mbf{\mu}_{y_i} \right|\right|_2^2 $$Given the scalar function:
$$ \mathcal{L}(\mbf{\mu};\mbf{D}) = \sum_{i=1}^N \left( {x}_i - {\mu} \right)^2 $$Write a proof sketch that ${\mu}$ as the average of $\mbf{D}\doteq\{{x}_i\}_{i=1}^N$ is a critical point of the function. (It is also a global minimum)
Given the function vector to scalar function, where the assignments $\bar{y}$ are given somehow (they cannot "move"):
$$ \mathcal{L}(\mbf{\mu},\bar{y};\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \boldsymbol{\mu}_{\bar{y}_i} \right|\right|_2^2 $$Write a proof sketch that $\boldsymbol{\mu}$ as the average of the vectors $\mbf{D}\doteq\{\mbf{x}_i\}_{i=1}^N$ is a critical point of the function. (It is also a global minimum)
Hint: set the derivative/gradient of mu to zero and solve; for 2b) pay attention to how the L2 norm is defined
Objective and Motivation: The goal of unsupervised learning is to find hidden patterns in unlabeled data.
\begin{equation} \underbrace{\{\mathbf{x}_i\}_{i=1}^N}_{\text{known}} \sim \underbrace{\mathcal{D}}_{\text{unknown}} \end{equation}Unlike in supervised learning, any data point is not paired with a label.
As you can see the unsupervised learning problem is ill-posed (which hidden patterns?) and in principle more difficult than supervised learning.
Unsupervised learning can be thought of as "finding structure" in the data.
We use log because:
log and exp and $\prod$ will change to $\sum$ because of log propertiesargmax, not maxor
$$ \boldsymbol{\theta}^* = \arg\max_{\boldsymbol{\theta}} \sum_{i=1}^N\ln \big( L(\boldsymbol{\mu}, \boldsymbol{\Sigma};\mbf{x}_i)\big) $$Objective and Motivation: The goal of unsupervised learning is to find hidden patterns in unlabeled data.
\begin{equation} \underbrace{\{\mathbf{x}_i\}_{i=1}^N}_{\text{known}} \sim \underbrace{\mathcal{D}}_{\text{unknown}} \end{equation}Unlike in supervised learning, any data points is not paired with a label.
As you can see the unsupervised learning problem is ill-posed (which hidden patterns?) and in principle more difficult than supervised learning.
Unsupervised learning can be thought of as "finding structure" in the data.
We could decide to model it with a normalized histogram
In practice, the histogram technique can be useful for obtaining a quick visualization of data in one or two dimensions but is unsuited to most density estimation applications
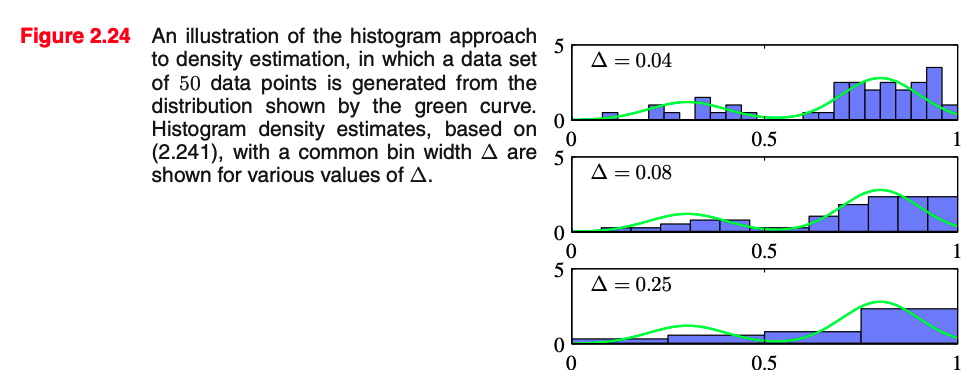

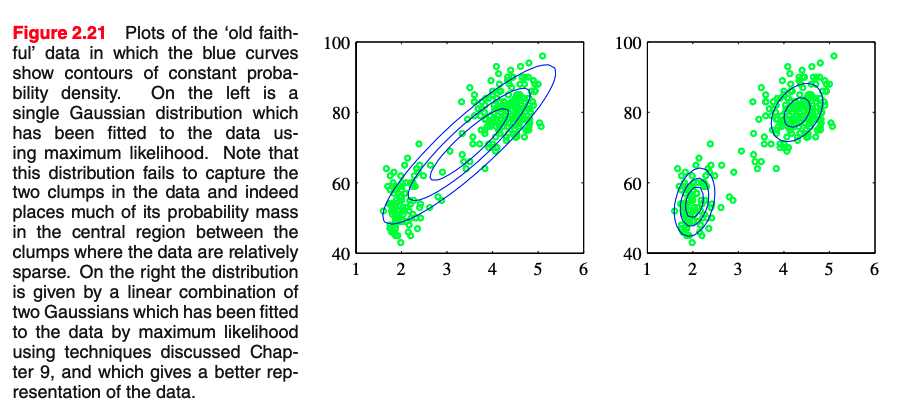
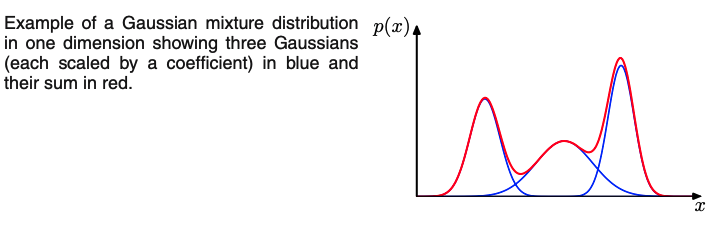
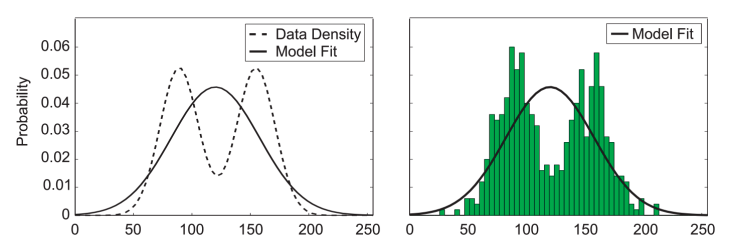
[Slide credit: K. Kutulakos]
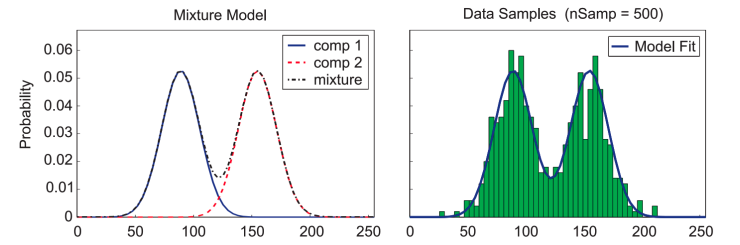
[Slide credit: K. Kutulakos]
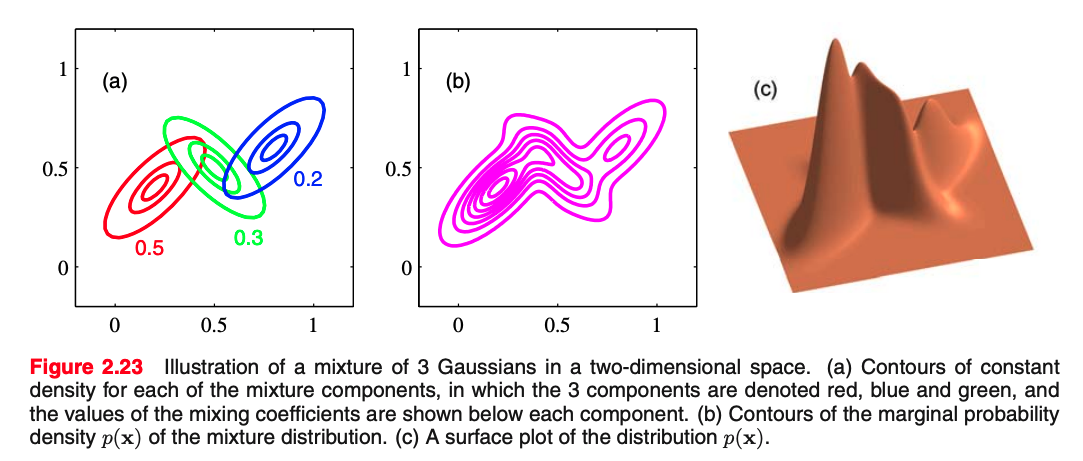
From the sum and product rules, we can express the data density $p(\mbf{x})$ as the marginal density where the marginalization is over the mode/Gaussians we have: $$ p(\mathbf{x})=\sum_{k=1}^{K} \underbrace{p(k)}_{\text{select mode}} \underbrace{p(\mathbf{x} \mid k)}_{\text{gen. point}} $$ which is equivalent to MoG equation, in which we can view $\pi_{k}=p(k)$ as the prior probability of picking the $k^{\text {th }}$ component, and the density $\mathcal{N}\left(\mathbf{x} \mid \boldsymbol{\mu}_{k}, \boldsymbol{\Sigma}_{k}\right)=p(\mathbf{x} \mid k)$ as the probability of $\mathbf{x}$ conditioned on $k$.
How can we generate points assuming we have the parameters of the model?
How can we generate points assuming we have a model with $3$ Gaussian where mixing coefficients are $\pi=[0.5, 0.3, 0.2]$;
A possible algorithm is:
A possible algorithm is:
import matplotlib.pyplot as plt
np.random.seed(0)
def inverse_sampling(pmf):
# import random
# random.choices(range(1,k), pmf, k=1000)
return np.argmin((np.random.rand(1)[:, None] > pmf.cumsum()), axis=1)
################### model params #######################
N_samples = 1000 # we sample this amount of points
z_to_gaussian = {0: ([0, 0], [[0.5, 0], [0, 0.15]], 'red'),
1: ([3, 2], [[1.25, 0], [0, 0.75]], 'green'),
2: ([-2, -2], [[1, -0.74], [-0.74, 1]], 'blue'),
}
mixing = np.array([0.05, 0.05, 0.9]) # mixing coefficients
########################################################
for _ in range(0, N_samples): # for each sample
z = inverse_sampling(mixing) # sample k using mixing coefficients
*normal, color = z_to_gaussian[z[0]] # now sample the data from N_k
x, y = np.random.multivariate_normal(
*normal, 1).T # sample 1 point at a time for clarity
plt.plot(x, y, '.', color=color)
plt.title('p(x,z)')
plt.axis('equal')
plt.show()
import matplotlib.pyplot as plt
np.random.seed(0)
def inverse_sampling(pmf):
return np.argmin((np.random.rand(1)[:, None] > pmf.cumsum()), axis=1)
################### model params #######################
N_samples = 1000 # we sample this amount of points
z_to_gaussian = {0: ([0, 0], [[0.5, 0], [0, 0.15]], 'red'),
1: ([3, 2], [[1.25, 0], [0, 0.75]], 'green'),
2: ([-2, -2], [[1, -0.74], [-0.74, 1]], 'blue'),
}
mixing = np.array([0.2, 0.5, 0.3]) # mixing coefficients
########################################################
for _ in range(0, N_samples): # for each sample
z = inverse_sampling(mixing) # sample k using mixing coefficients
*normal, color = z_to_gaussian[z[0]] # now sample the data from N_k
x, y = np.random.multivariate_normal(
*normal, 1).T # sample 1 point at time for clarity
plt.plot(x, y, '.', color='black')
plt.axis('equal')
plt.title('p(x)')
plt.show()
from which Gaussian is this point sampled? Alas, we do not have the answer to this question.Let's assume $k=3$ Gaussian and $X=\{\mbf{x}_1,\ldots,\mbf{x}_N \}$ and $Y=\{{y}_1,\ldots,{y}_N \}$
where $Y$ denotes an index from 0 to k-1 that selects the Gaussian for that point.
If we had, we could do:
$\bbox[yellow, 5px]{Problem:}$ we cannot assume to know the association of the points to Gaussians. (i..e we do not know $Y$).
We can do MLE on a Mixture of Gaussians:
Problems:
A (trivial) way of thinking about a data set of points to fit a parametric model or to estimate a density is to:
This is related to the idea of overfitting the data (more on overfitting in supervised learning).

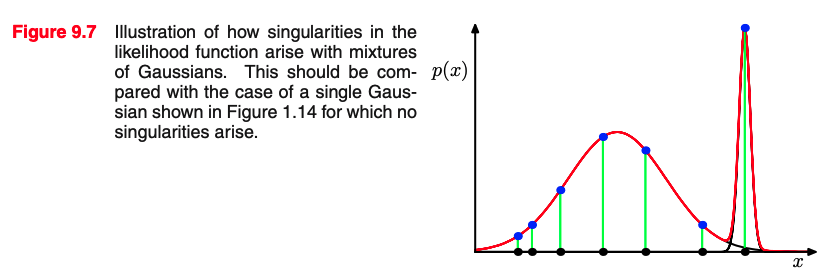
We can do MLE on a Mixture of Gaussians:
Assumptions:
Variables that are always unobserved are called latent variables, or sometimes hidden variables
We may want to intentionally introduce latent variables to model complex dependencies between variables – this can simplify the model
In a mixture model, the identity of the component that generated a given data point is a latent variable
It is made of two steps:
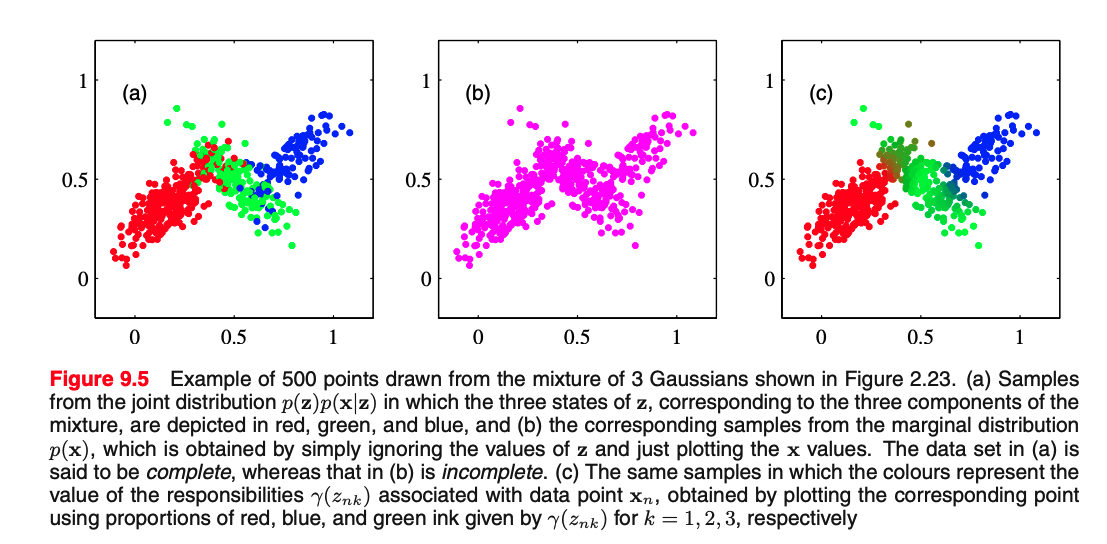
Given a data point, what is the probability comes from Gaussian $\mbf{z}==k$?
Given a data point, what is the probability comes from Gaussian $\mbf{z}==k$?
$$ \gamma_{k} = p(\bz=k | \bx) = \frac{p(\bx | \bz=k) p(\bz=k)}{p(\bx)} = \frac{p(\bx | \bz=k) p(\bz=k)}{\sum_j^K p(\bx | \bz=j) p(\bz=j)} $$[0.8, 0.1, 0.1] for 3 Gaussians
import matplotlib.pyplot as plt
np.random.seed(0)
def inverse_sampling(pmf):
return np.argmin((np.random.rand(1)[:, None] > pmf.cumsum()), axis=1)
################### model params #######################
N_samples = 1000 # we sample this amount of points
z_to_gaussian = {0: ([0, 0], [[0.5, 0], [0, 0.15]], 'red'),
1: ([3, 2], [[1.25, 0], [0, 0.75]], 'green'),
2: ([-2, -2], [[1, -0.74], [-0.74, 1]], 'blue'),
}
mixing = np.array([0.2, 0.5, 0.3]) # mixing coefficients
########################################################
for _ in range(0, N_samples): # for each sample
z = inverse_sampling(mixing) # sample k using mixing coefficients
*normal, color = z_to_gaussian[z[0]] # now sample the data from N_k
x, y = np.random.multivariate_normal(
*normal, 1).T # sample 1 point at a time for clarity
plt.plot(x, y, '.', color='black')
plt.axis('equal')
plt.title('p(x)')
plt.show()
from sklearn.mixture import GaussianMixture
# X is 2xN
gmm = GaussianMixture(n_components=3, random_state=0).fit(X.T)
# >>> gm.means_ #Dxk
assignments = gmm.predict(X.T)
z_to_col = {0: 'red', 1: 'green', 2: 'blue'}
_ = plt.scatter(X[0, ...], X[1, ...], color=[z_to_col[y] for y in assignments])
_ = plt.scatter(*gmm.means_.T,
s=250,
marker='*',
c='black',
label='centroids')
from matplotlib.colors import LogNorm
fig = plt.figure(figsize=(10, 10))
# display predicted scores by the model as a contour plot
x = np.linspace(-4.0, 6.0)
y = np.linspace(-4.0, 6.0)
X1, Y1 = np.meshgrid(x, y)
XX = np.array([X1.ravel(), Y1.ravel()]).T
Z = -gmm.score_samples(XX) # negative log likelihood
Z = Z.reshape(X1.shape)
CS = plt.contourf(
X1, Y1, Z, cmap='hot'
)
plt.grid(False)
CB = plt.colorbar(CS, shrink=0.8, extend="both")
plt.scatter(X[0, ...], X[1, ...])
plt.title("Negative log-likelihood predicted by a GMM")
plt.axis("tight")
plt.show()
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import numpy as np
# Generate some data
from sklearn.datasets import make_blobs
X, y_true = make_blobs(n_samples=400, centers=4,
cluster_std=0.60, random_state=0)
X = X[:, ::-1] # flip axes for better plotting
# Plot the data with K Means Labels
from sklearn.cluster import KMeans
kmeans = KMeans(4, random_state=0)
labels = kmeans.fit(X).predict(X)
plt.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis');
from sklearn.cluster import KMeans
from scipy.spatial.distance import cdist
def plot_kmeans(kmeans, X, n_clusters=4, rseed=0, ax=None):
labels = kmeans.fit_predict(X)
# plot the input data
ax = ax or plt.gca()
ax.axis('equal')
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis', zorder=2)
# plot the representation of the KMeans model
centers = kmeans.cluster_centers_
radii = [cdist(X[labels == i], [center]).max()
for i, center in enumerate(centers)]
for c, r in zip(centers, radii):
ax.add_patch(plt.Circle(c, r, fc='#CCCCCC', lw=3, alpha=0.5, zorder=1))
kmeans = KMeans(n_clusters=4, random_state=0)
plot_kmeans(kmeans, X)
rng = np.random.RandomState(13)
X_stretched = np.dot(X, rng.randn(2, 2))
kmeans = KMeans(n_clusters=4, random_state=0)
plot_kmeans(kmeans, X_stretched)
from matplotlib.patches import Ellipse
def draw_ellipse(position, covariance, ax=None, **kwargs):
"""Draw an ellipse with a given position and covariance"""
ax = ax or plt.gca()
# Convert covariance to principal axes
if covariance.shape == (2, 2):
U, s, Vt = np.linalg.svd(covariance)
angle = np.degrees(np.arctan2(U[1, 0], U[0, 0]))
width, height = 2 * np.sqrt(s)
else:
angle = 0
width, height = 2 * np.sqrt(covariance)
# Draw the Ellipse
for nsig in range(1, 4):
ax.add_patch(Ellipse(position, nsig * width, nsig * height,
angle, **kwargs))
def plot_gmm(gmm, X, label=True, ax=None):
ax = ax or plt.gca()
labels = gmm.fit(X).predict(X)
if label:
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis', zorder=2)
else:
ax.scatter(X[:, 0], X[:, 1], s=40, zorder=2)
ax.axis('equal')
w_factor = 0.2 / gmm.weights_.max()
for pos, covar, w in zip(gmm.means_, gmm.covariances_, gmm.weights_):
draw_ellipse(pos, covar, alpha=w * w_factor)
from sklearn.mixture import GaussianMixture
gmm = GaussianMixture(n_components=4).fit(X)
labels = gmm.predict(X)
plt.scatter(X[:, 0], X[:, 1], c=labels, s=40, cmap='viridis');
gmm = GaussianMixture(n_components=4, random_state=42)
plot_gmm(gmm, X)
gmm = GaussianMixture(n_components=4, covariance_type='full', random_state=42)
plot_gmm(gmm, X_stretched)
GMM can have assumptions even on the shape of the Gaussian based on the covariance matrix.
Remember:
$\bsigma$ types:
full: each component has its own general covariance matrix (more parameters)tied: all components share the same general covariance matrix.diag each component has its own diagonal covariance matrix.spherical: each component has its own single variance. (less params)from sklearn.datasets import make_moons
Xmoon, ymoon = make_moons(200, noise=.05, random_state=0)
plt.scatter(Xmoon[:, 0], Xmoon[:, 1]);
gmm2 = GaussianMixture(n_components=2, covariance_type='full', random_state=0)
plot_gmm(gmm2, Xmoon)
gmm16 = GaussianMixture(n_components=8, covariance_type='full', random_state=0)
plot_gmm(gmm16, Xmoon, label=False)
Note that sample means generate
Xnew, _ = gmm16.sample(4000)
plot_gmm(gmm16, Xmoon, label=True)
plt.scatter(Xnew[:, 0], Xnew[:, 1], color='red')
Xnew, _ = gmm16.sample(40000)
loglike = gmm16.score_samples(Xnew)
plt.scatter(Xnew[:, 0], Xnew[:, 1], c=loglike, cmap='jet')
n_components = np.arange(1, 21)
models = [GaussianMixture(n, covariance_type='full', random_state=0).fit(Xmoon)
for n in n_components]
plt.plot(n_components, [m.bic(Xmoon) for m in models], label='BIC')
plt.plot(n_components, [m.aic(Xmoon) for m in models], label='AIC')
plt.legend(loc='best')
plt.xlabel('n_components');
from sklearn.datasets import load_digits
digits = load_digits()
digits.data.shape
def plot_digits(data):
fig, ax = plt.subplots(10, 10, figsize=(8, 8),
subplot_kw=dict(xticks=[], yticks=[]))
fig.subplots_adjust(hspace=0.05, wspace=0.05)
for i, axi in enumerate(ax.flat):
im = axi.imshow(data[i].reshape(8, 8), cmap='gray')
im.set_clim(0, 16)
plot_digits(digits.data)
from sklearn.decomposition import PCA
pca = PCA(0.99, whiten=False)
data = pca.fit_transform(digits.data)
data.shape
n_components = np.arange(1, 40, 1)
models = [GaussianMixture(n, covariance_type='diag', random_state=0)
for n in n_components]
aics = [model.fit(data).bic(data) for model in models]
plt.plot(n_components, aics);
idx = (n_components==10).nonzero()[0][0]
model = models[idx]
data_new, y_new = model.sample(100)
data_new.shape
digits_new = pca.inverse_transform(data_new)
plot_digits(digits_new)
plt.bar(*np.unique(digits.target, return_counts=True));
plt.bar(range(10),model.weights_);
plt.bar(range(model.means_[0].shape[0]),model.means_[0]);
plt.bar(range(model.means_[0].shape[0]),model.covariances_[0]);
digits.target with K=10