

The geometric view of linear algebra gives an intuitive way to interpret a fundamental quantity known as the determinant. Consider the grid image from before, but now with a highlighted region below.
Look at the highlighted square. This is a square with edges given by $(0, 1)$ and $(1, 0)$ and thus it has area one. After $\mathbf{A}$ transforms this square, we see that it becomes a parallelogram. There is no reason this parallelogram should have the same area that we started with, and indeed in the specific case shown here of
$$ \mathbf{A} = \begin{bmatrix} 1 & 2 \\ -1 & 3 \end{bmatrix}, $$it is an exercise in coordinate geometry to compute the area of this parallelogram and obtain that the area is $5$.
In general, if we have a matrix
$$ \mathbf{A} = \begin{bmatrix} a & b \\ c & d \end{bmatrix}, $$we can see with some computation that the area of the resulting parallelogram is $ad-bc$. This area is referred to as the determinant.
Sanity Check: We cannot apply Pythagoras' Theorem to compute area because axes are not aligned anymore.
The picture is misleading since the axes are CLOSED to be aligned.
The angle between $[1, -1]$ and $[2,3]$ is 101.30993247402021 ```python import numpy as np; X = np.array([[1, -1], [2, 3]]);thetarad = angle(X[0,:],X[1,:]);theta = thetarad*180/np.pi; print(theta)}}
The angle between $[1, 0]$ and $[0, 1]$ is 90.
```python
is_basis = False # toggle this to compare the angles
X = np.array([[1, 0], [0, 1]]) if is_basis else np.array([[1, -1], [2, 3]])
theta_rad = angle(*X)
theta = theta_rad*180/np.pi
print(theta)is_basis = False # toggle this to compare the angles
X = np.array([[1, 0], [0, 1]]) if is_basis else np.array([[1, -1], [2, 3]])
theta_rad = angle(*X)
theta = theta_rad*180/np.pi
print(theta)
101.30993247402021
Preserve angles and distances
$$A = \left[\begin{array}{cc}\cos\theta ~~~ -\sin\theta \\\sin\theta ~~~ \cos\theta\end{array} \right]$$This is easily derived by noting that
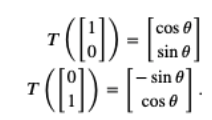
ang = np.pi/4
A = np.array([[np.cos(ang), -np.sin(ang)],
[np.sin(ang),np.cos(ang)]])
print(A)
Xd, Yd = linear_map(A, Xs, Ys)
fig, axs = plt.subplots(1,2)
fig.suptitle('Rotation')
plot_grid(Xs,Ys,axs[0])
plot_grid(Xd,Yd,axs[1])
[[ 0.70710678 -0.70710678] [ 0.70710678 0.70710678]]
Preserve angles and RATIO between distances
$$A = \left[\begin{array}{cc}s_x\cos\theta ~~~ -\sin\theta \\\sin\theta ~~~ s_y\cos\theta\end{array} \right]$$This is easily derived by noting that
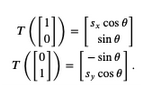
ang, scale, clip = np.pi/4, 2, True
A = np.array([[scale*np.cos(ang), -np.sin(ang)],
[np.sin(ang), scale*np.cos(ang)]])
print(A)
Xd, Yd = linear_map(A, Xs, Ys)
fig, axs = plt.subplots(1, 2)
fig.suptitle('Rotation')
plot_grid(Xs, Ys, axs[0])
plot_grid(Xd, Yd, axs[1])
if clip:
plt.xlim(-nX, nX)
plt.ylim(-nY, nY)
[[ 1.41421356 -0.70710678] [ 0.70710678 1.41421356]]
Preserve parallelism (but NOT angles!)
$$A = \left[\begin{array}{cc}s_x\cos\theta ~~~ -c_x\sin\theta \\c_y\sin\theta ~~~ s_y\cos\theta\end{array} \right]$$This is easily derived by noting that
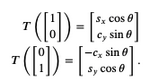
ang, scale, cx_shear, cy_shear,clip = np.pi/4, 2, 8, 2, False
A = np.array([[scale*np.cos(ang), -cx_shear*np.sin(ang)],
[cy_shear*np.sin(ang), scale*np.cos(ang)]])
print(A)
Xd, Yd = linear_map(A, Xs, Ys)
fig, axs = plt.subplots(1, 2)
fig.suptitle('Rotation')
plot_grid(Xs, Ys, axs[0])
plot_grid(Xd, Yd, axs[1])
if clip:
plt.xlim(-nX, nX)
plt.ylim(-nY, nY)
[[ 1.41421356 -5.65685425] [ 1.41421356 1.41421356]]
Hyperplane: a generalization to higher dimensions of a line ($D=2$) or of a plane ($D=3$). In a $d$-dimensional vector space, a hyperplane has $d-1$ dimensions and divides the space into two half-spaces.
$$ \mathbf{w}\mathbf{x} + \mathbf{b} = 0 $$where $\mathbf{w}$ is a vector normal to the hyperplane and $\mathbf{b}$ is an offset
Hyperplane: a generalization to higher dimensions of a line ($D=2$) or of a plane ($D=3$). In an $d$-dimensional vector space, a hyperplane has $d-1$ dimensions and divides the space into two half-spaces.
$$ \mathbf{w}\mathbf{x} + \mathbf{b} = 0$$where $\mathbf{w}$ is a vector normal to the hyperplane and $\mathbf{b}$ is an offset
Suppose that we have two vectors $\mathbf{v}$ and a column vector $\mathbf{w}=[2,1]^\top$.
We want to project $\mathbf{v}$ onto $\mathbf{w}$ or better project v onto the subspace (line in this case) of $\mathbf{w}$.
Recalling trigonometry, we see the formula $\|\mathbf{v}\|\cos(\theta)$ is the length of the projection of the vector $\mathbf{v}$ onto the direction of $\mathbf{w}$
Defining a unit vector $\mathbf{\hat{w}}=\frac{\mathbf{w}}{||\mathbf{w}||}$ we have: $$\mathbb{P}_\mathbf{w}(\mathbf{v}) = \underbrace{\mathbf{\hat{w}}}_{\text{direction}} \left(\underbrace{\mathbf{\hat{w}}^T \mathbf{v}}_{\text{length}}\right)$$
Suppose that we have a matrix $A$ with the following entries:
$$ \mathbf{A} = \begin{bmatrix} 2 & 0 \\ 0 & -1 \end{bmatrix}. $$If we apply $A$ to any vector $\mathbf{v} = [x, y]^\top$, we obtain a vector $\mathbf{A}\mathbf{v} = [2x, -y]^\top$. This has an intuitive interpretation: stretch the vector to be twice as wide in the $x$-direction, and then flip it in the $y$-direction.
However, there are some vectors for which something remains unchanged.
Namely $[1, 0]^\top$ gets sent to $[2, 0]^\top$ and $[0, 1]^\top$ gets sent to $[0, -1]^\top$.
These vectors are still in the same line, and the only modification is that the matrix stretches them by a factor of $2$ and $-1$ respectively. We call such vectors eigenvectors and the factor they are stretched by eigenvalues.
In general, if we can find a number $\lambda$ and a vector $\mathbf{v}$ such that
$$ \underbrace{\mathbf{A}}_{\text{known}}\mathbf{v} = \underbrace{\lambda\mathbf{v}}_{\text{unknown}} $$We say that $\mathbf{v}$ is an eigenvector for $A$ and $\lambda$ is an eigenvalue.
For the equation to happen, we see that $(\mathbf{A} - \lambda \mathbf{I})$ must compress some direction down to zero, hence it is not invertible, and thus the determinant is zero. Thus, we can find the eigenvalues by finding for what $\lambda$ is $\det(\mathbf{A}-\lambda \mathbf{I}) = 0$. Once we find the eigenvalues, we can solve $\mathbf{A}\mathbf{v} = \lambda \mathbf{v}$ to find the associated eigenvector(s).
Let's see this with a more challenging matrix
$$ \mathbf{A} = \begin{bmatrix} 2 & 1\\ 2 & 3 \end{bmatrix}. $$If we consider $\det(\mathbf{A}-\lambda \mathbf{I}) = 0$, we see this is equivalent to the polynomial equation $0 = (2-\lambda)(3-\lambda)-2 = (4-\lambda)(1-\lambda)$. Thus, two eigenvalues are $4$ and $1$. To find the associated vectors, we then need to solve
$$ \begin{bmatrix} 2 & 1\\ 2 & 3 \end{bmatrix}\begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}1x \\ 1y\end{bmatrix} \; \text{and} \; \begin{bmatrix} 2 & 1\\ 2 & 3 \end{bmatrix}\begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}4x \\ 4y\end{bmatrix} . $$We can solve this with the vectors $[1, -1]^\top$ and $[1, 2]^\top$ respectively.
eigs, eigvects = np.linalg.eig(
np.array([[2, 1],
[2, 3]]))
print(f'Eigen val:\n {eigs}', end='\n\n')
print(f'EigenVect vect:\n {eigvects}')
# Note that `numpy` normalizes the eigenvectors to be of length one,
# whereas we took ours to be of arbitrary length.
# Additionally, the choice of the sign is arbitrary.
# However, the vectors computed are parallel
# to the ones we found by hand with the same eigenvalues.
#help(np.linalg.eig)
Eigen val: [1. 4.] EigenVect vect: [[-0.70710678 -0.4472136 ] [ 0.70710678 -0.89442719]]
fig_dim = 10
### Unit Sphere
# Parametric sphere
t = np.linspace(0, 2*np.pi, 100) # seems a circle but it is not (move the resolution to 10)
Xs = np.cos(t)
Ys = np.sin(t)
# Plot
fig, ax = plt.subplots()
fig.set_figheight(fig_dim)
fig.set_figwidth(fig_dim)
plot_points(ax, Xs, Ys, col='blue')
plt.show()
fig_dim = 7
### Unit Sphere
# Parametric sphere
t = np.linspace(0, 2*np.pi, 100) # seems a circle but it is not (move the resolution to 10)
Xs = np.cos(t)
Ys = np.sin(t)
# Plot
fig, ax = plt.subplots()
fig.set_figheight(fig_dim)
fig.set_figwidth(fig_dim)
plot_points(ax, Xs, Ys, col='blue')
plt.show()
We can do it without using for loop (we used it in the previous lecture)
A@PFor now, assume:
A is square# Define a transform
limit = 2.5
# Define a transformation
# A = np.array([[1.5, 1],
# [1, 1.5]])
A = np.array([[1, -1],
[-1, 2]])
print(f'Transformation is\n {A}')
Transformation is [[ 1 -1] [-1 2]]
# Map points on the surface
Xd, Yd = linear_map(A, Xs, Ys)
# let's map the basis
Xu_d, Yu_d = linear_map(A, np.array([1,0]), np.array([0,1]))
units_d = np.stack((Xu_d, Yu_d),axis=1)
# Compute the eigenvalues and eigenvectors
eigVals, eigVecs = np.linalg.eig(A)
print('eigVals', eigVals, 'eigVecs', eigVecs, sep='\n'*2)
eigVals [0.38196601 2.61803399] eigVecs [[-0.85065081 0.52573111] [-0.52573111 -0.85065081]]
# Plot src points and destination points
fig, ax = plt.subplots(1,1)
fig.set_figheight(fig_dim)
fig.set_figwidth(fig_dim)
# Plot src
plot_points(ax, Xs, Ys, col='blue')
# Plot destination
plot_points(ax, Xd, Yd, col='green')
ax.set_aspect('equal')
ax.set_xlim(-limit,limit)
ax.set_ylim(-limit,limit)
# Plot normalized eigenvectors (unit 1)
plotVectors(ax, [eigVecs[:,0], eigVecs[:,1]],
cols=['red']*2)
# Plot unnormalized eigenvectors (unit 1)
# we have to multiply back to its own eig val
plotVectors(ax, [eigVals[0]*eigVecs[:,0], eigVals[1]*eigVecs[:,1]],
cols=['pink']*2)
plotVectors(ax, [Xu_d, Yu_d ],
cols=['gray']*2)
# Plot src points and destination points
fig, ax = plt.subplots(1,1)
fig.set_figheight(fig_dim)
fig.set_figwidth(fig_dim)
# Plot src
plot_points(ax, Xs, Ys, col='blue')
# Plot destination
plot_points(ax, Xd, Yd, col='green')
ax.set_aspect('equal')
ax.set_xlim(-limit,limit)
ax.set_ylim(-limit,limit)
# Plot normalized eigenvectors (unit 1)
plotVectors(ax, [eigVecs[:,0], eigVecs[:,1]],
cols=['red']*2)
# Plot unnormalized eigenvectors (unit 1)
# we have to multiply back to its own eig val
plotVectors(ax, [eigVals[0]*eigVecs[:,0], eigVals[1]*eigVecs[:,1]],
cols=['pink']*2)
plotVectors(ax, [Xu_d, Yu_d ],
cols=['gray']*2)
np.prod(eigVals) = NameError: name 'np' is not defined
be the matrix where the columns are the eigenvectors of the matrix $\mathbf{A}$. Let
$$ \boldsymbol{\Sigma} = \begin{bmatrix} 1 & 0 \\ 0 & 4 \end{bmatrix}, $$be the matrix with the associated eigenvalues on the diagonal. Then the definition of eigenvalues and eigenvectors tells us that
$$ \mathbf{A}\mathbf{W} =\mathbf{W} \boldsymbol{\Sigma} . $$The matrix $W$ is invertible, so we may multiply both sides by $W^{-1}$ on the right, we see that we may write
$$\mathbf{A} = \mathbf{W} \boldsymbol{\Sigma} \mathbf{W}^{-1}.$$if $\mathbf{W}$ is invertible
One nice thing about eigendecompositions is that we can write many operations we usually encounter cleanly in terms of the eigendecomposition. As a first example, consider:
$$ \mathbf{A}^n = \overbrace{\mathbf{A}\cdots \mathbf{A}}^{\text{$n$ times}} = \overbrace{(\mathbf{W}\boldsymbol{\Sigma} \mathbf{W}^{-1})\cdots(\mathbf{W}\boldsymbol{\Sigma} \mathbf{W}^{-1})}^{\text{$n$ times}} = \mathbf{W}\overbrace{\boldsymbol{\Sigma}\cdots\boldsymbol{\Sigma}}^{\text{$n$ times}}\mathbf{W}^{-1} = \mathbf{W}\boldsymbol{\Sigma}^n \mathbf{W}^{-1}. $$This tells us that for any positive power of a matrix, the eigendecomposition is obtained by just raising the eigenvalues to the same power. The same can be shown for negative powers, so if we want to invert a matrix we need only consider
$$ \mathbf{A}^{-1} = \mathbf{W}\boldsymbol{\Sigma}^{-1} \mathbf{W}^{-1}, $$or in other words, just invert each eigenvalue.
Informal: Given a matrix $\mathbf{A}$ squared and symmetric, $\mathbf{A} \in \mathbb{R}^{d\times d}$ and $\mathbf{A} = \mathbf{A}^T$ then:
| Method | $\mathbf{A}$ | Decompostion |
|---|---|---|
SVD | any | $\mathbf{A} = \mathbf{U}\mathbf{\Sigma}\mathbf{V}^{\top}$ Eig | square | $\mathbf{A} = \mathbf{U}\mathbf{\Sigma}\mathbf{U}^{-1}$ Eig=SVD | square/sym | $\mathbf{A} = \mathbf{U}\mathbf{\Sigma}\mathbf{U}^{\top}$
| Method | Step 1 | Step 2 | Step 3 | |
|---|---|---|---|---|
geometry | |rotate| scale\reflect| rotate SVD | | $\mathbf{V}^{\top}$ | $\mathbf{\Sigma}$ | $\mathbf{U}$ geometry | |rotate| scale\reflect| rotate Eig | | $\mathbf{U}^{-1}$ | $\mathbf{\Sigma}$ | $\mathbf{U}$
print('A', A, sep='\n\n')
Sigma, U = np.diag(eigVals), eigVecs
print('Sigma', Sigma, 'U', U, sep='\n\n')
A [[ 1 -1] [-1 2]] Sigma [[0.38196601 0. ] [0. 2.61803399]] U [[-0.85065081 0.52573111] [-0.52573111 -0.85065081]]
# Plot src points and destination points
fig, axes = plt.subplots(1, 4)
fig.set_figheight(20)
fig.set_figwidth(20)
# apply [Id, Uinv, Sigma, U]
Trans = [np.diag((1, 1)), U.T, Sigma, U]
titles = ['Init', 'Step 1 Uinv', 'Step 2 Sigma', 'Step 3 U']
X_orig = np.array([1, 0])
Y_orig = np.array([0, 1])
Xss = np.copy(Xs)
Yss = np.copy(Ys)
for count, (ax, T, title) in enumerate(zip(axes, Trans, titles)):
ax.set_xlim(-limit, limit)
ax.set_ylim(-limit, limit)
ax.set_aspect('equal')
ax.set_title(title)
Xss, Yss = linear_map(T, Xss, Yss)
X_orig, Y_orig = linear_map(T, X_orig, Y_orig)
if count == 0:
plot_points(ax, Xs, Ys, col='blue')
unit_v = np.stack((X_orig, Y_orig), axis=1)
plotVectors(ax, unit_v, ['blue', 'red'])
plot_points(ax, Xss, Yss, col='green')
# Plot src points and destination points
fig, axes = plt.subplots(1, 4)
fig.set_figheight(20)
fig.set_figwidth(20)
# apply [Id, Uinv, Sigma, U]
Trans = [np.diag((1, 1)), U.T, Sigma, U]
titles = ['Init', 'Step 1 Uinv', 'Step 2 Sigma', 'Step 3 U']
X_orig = np.array([1, 0])
Y_orig = np.array([0, 1])
Xss = np.copy(Xs)
Yss = np.copy(Ys)
for count, (ax, T, title) in enumerate(zip(axes, Trans, titles)):
ax.set_xlim(-limit, limit)
ax.set_ylim(-limit, limit)
ax.set_aspect('equal')
ax.set_title(title)
Xss, Yss = linear_map(T, Xss, Yss)
X_orig, Y_orig = linear_map(T, X_orig, Y_orig)
if count == 0:
plot_points(ax, Xs, Ys, col='blue')
unit_v = np.stack((X_orig, Y_orig), axis=1)
plotVectors(ax, unit_v, ['blue', 'red'])
plot_points(ax, Xss, Yss, col='green')
Source: Wikipedia

########### We know the generative model of the data D ############
np.random.seed(0) # fixing the seed
n_samples=100; cov = [[3, 3], [3, 4]]
# Assumes know the generative model of data
X = np.random.multivariate_normal(mean=[1, 1], cov=cov, size=n_samples)
###########################################################
print(f'num of points {X.shape[0]} in dimension {X.shape[1]}')
fig = plt.figure(figsize=(8,8))
plt.scatter(*X.T)
plt.scatter(0, 0, c='r', marker='o')
plt.ylabel('Enjoyment')
_ = plt.xlabel('Skills')
num of points 100 in dimension 2
NameError: name 'X' is not defined
then the covariance matrix needs to be $ \in \mathbb{R}^{D\times D}$: $$\mbf{C} = \frac{1}{N} (\mathbf{X}-\boldsymbol{\mu})^T(\mathbf{X}-\boldsymbol{\mu}) $$
NameError: name 'X' is not defined
then the covariance matrix needs to be $ \in \mathbb{R}^{D\times D}$: $$\mbf{C} = \frac{1}{N} (\mathbf{X}-\boldsymbol{\mu})(\mathbf{X}-\boldsymbol{\mu})^T $$
After: $ \mathbf{X}^{\prime}$ is standardized
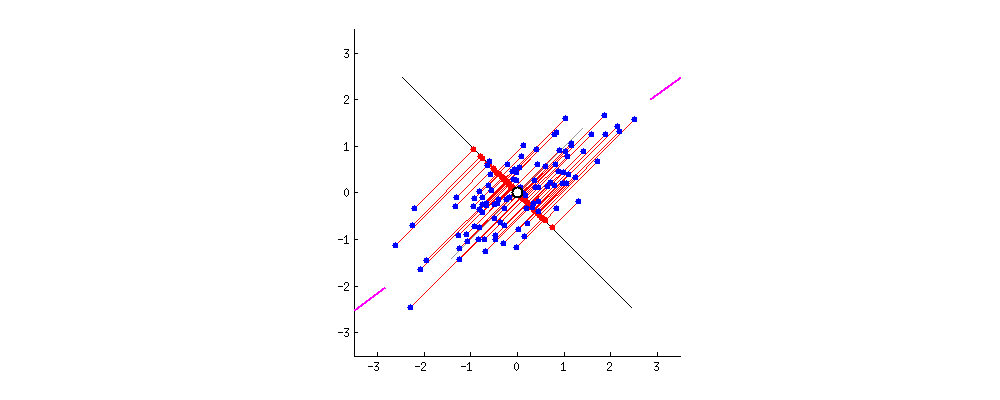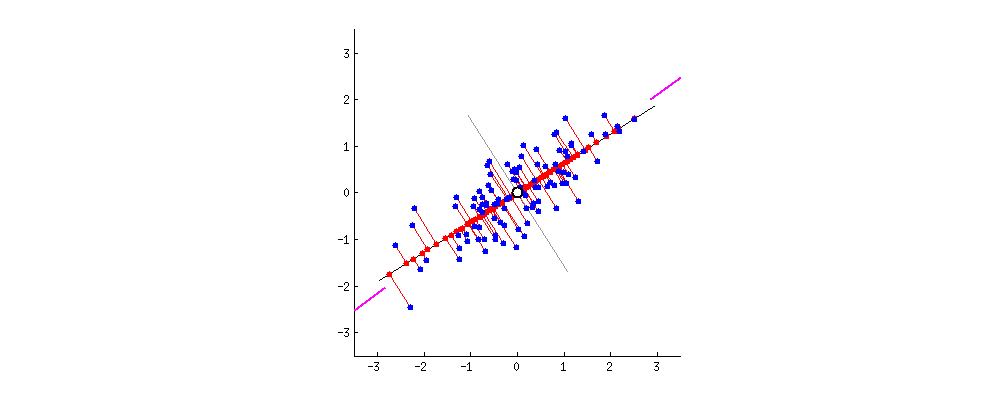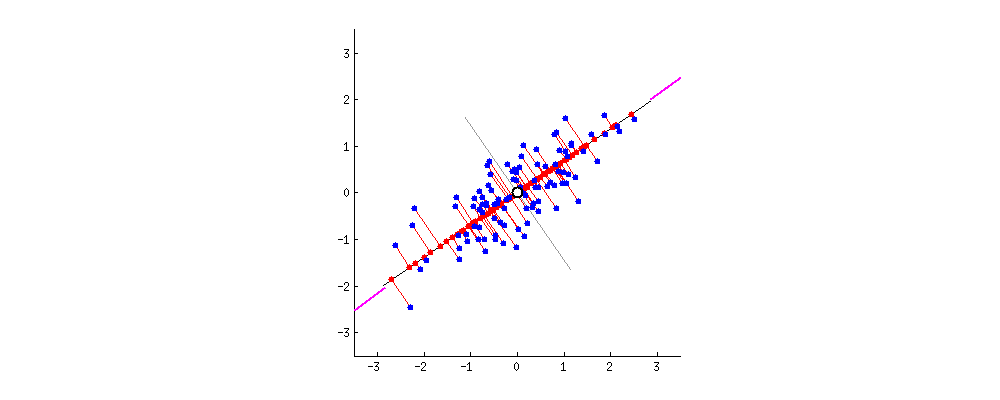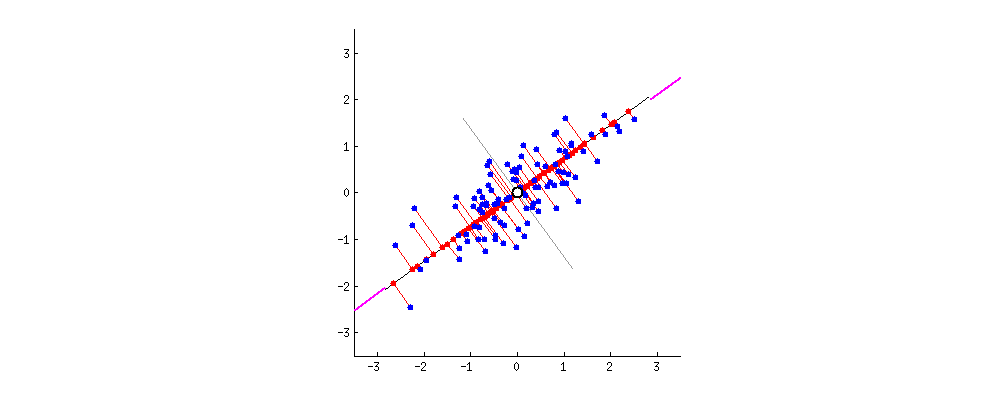 Taken from
Taken from
Can be used for compressing and reconstructing the data using U up to k components:
$$\underbrace{\mathbf{x}^T}_{\text{rec}} = \underbrace{\underbrace{\mathbf{U}_{|k}}_{k \mapsto D}\quad\underbrace{\underbrace{\mathbf{U}_{|k}^T}_{D \mapsto k}\quad\underbrace{\mathbf{x}^T}_{\text{orig}}}_{projection}}_{reconstruction}$$Note: *numpy is not guaranteed to return it ordered so you have to sort*
That is, keep $k$ components until:
$$\frac{\sum_i^k \lambda_i}{\sum_i^d \lambda_i} \leq 95\%$$print(f'num of points {X.shape[0]} in dimension {X.shape[1]}')
fig = plt.figure(figsize=(8,8))
plt.scatter(*X.T, alpha=0.3)
plt.scatter(0, 0, c='red')
plt.ylabel('Enjoyment')
plt.xlabel('Skills')
plt.axis('scaled')
plt.xlim(-6, 6)
plt.ylim(-6, 6)
num of points 100 in dimension 2
(-6.0, 6.0)
# standardize
center = X.mean(axis=0) #X shape is 100x2
std = X.std(axis=0)
Xp = (X-center)/std
fig = plt.scatter(*X.T, alpha=0.3)
plt.scatter(0, 0, c='red')
plt.quiver(0, 0, *center, angles='xy', scale_units='xy', scale=1)
plt.ylabel('Enjoyment')
plt.xlabel('Skills')
plt.axis('scaled')
plt.xlim(-3, 3)
_=plt.ylim(-3, 3)
#print(center)
fig = plt.figure(figsize=(8,8))
plt.scatter(*Xp.T, alpha=0.3)
plt.scatter(0, 0, c = 'red')
plt.ylabel('Enjoyment')
plt.xlabel('Skills')
plt.axis('scaled')
plt.xlim(-6, 6)
_=plt.ylim(-6, 6)
C = np.cov(X, rowvar=False)
Sigma, U = np.linalg.eig(C)
# If rowvar is True (default), then each row represents a variable,
# with observations in the columns. Otherwise, the relationship
# is transposed: each column represents a variable, while the
# rows contain observations.
C.shape
(2, 2)
Sigma.shape, U.shape, Sigma
((2,), (2, 2), array([0.47772418, 6.90110921]))
total_energy = Sigma.sum()
var_exp = Sigma/total_energy
cum_var_exp = np.cumsum(var_exp)
plt.bar(range(len(var_exp)), var_exp, alpha=0.5, align='center',
label='individual explained variance')
plt.step(range(len(var_exp)), cum_var_exp, where='mid',
label='cumulative explained variance')
plt.ylabel('Explained variance ratio')
plt.xlabel('Principal components')
plt.legend(loc='best')
plt.tight_layout()
plt.show()
Sigma = np.diag(Sigma)
Can be seen as reconstructing the data using U: $$\underbrace{\mathbf{x}^T}_{2 \times N} = \underbrace{\mathbf{U}}_{2\times 2}\quad\underbrace{\mathbf{U}^T}_{2\times 2}\quad\underbrace{\mathbf{x}^T}_{2\times N}$$
# Full projection
print(U.shape, U.T.shape, Xp.T.shape)
Xd = U @ U.T @ Xp.T # Our transformation
print(Xd.shape)
(2, 2) (2, 2) (2, 100) (2, 100)
print('Sigma', Sigma, 'U', U, sep='\n\n')
Sigma [[0.47772418 0. ] [0. 6.90110921]] U [[-0.76738982 -0.64118083] [ 0.64118083 -0.76738982]]
Utrunc = U[:,1].reshape(2,-1) # need reshape for matrix mul.
# note [:,1] selects the eigenvector with more energy (highest eigen value)
# if you have more of them you have to sort, with 2 it is easier it is either this or the other
print()
# Compressed projection
print('Full projection>', U.shape, U.T.shape, Xp.T.shape)
print('Compressed projection>', Utrunc.shape, Utrunc.T.shape, Xp.T.shape)
Xd = Utrunc.T @ Xp.T # Our transformation project down = Utrunc.T @ x.T;
print(Xd.shape)
Full projection> (2, 2) (2, 2) (2, 100) Compressed projection> (2, 1) (1, 2) (2, 100) (1, 100)
fig = plt.figure(figsize=(8,1))
plt.plot(Xd[0, ...].T, [0]*Xd.shape[1], 'o')
_ = plt.ylim(-0.001, 0.001)
fig = plt.figure(figsize=(8,8))
plt.scatter(*Xp.T, alpha=0.3)
plt.scatter(0, 0, c = 'red')
plt.ylabel('Enjoyment')
plt.xlabel('Skills')
plt.axis('scaled')
plt.xlim(-6, 6)
_=plt.ylim(-6, 6)
# Compressed projection and back-projected
print(U.shape, U.T.shape, Xp.T.shape)
Xd = Utrunc @ Utrunc.T @ Xp.T # Our transformation A = U @ Ut;
print(Xd.shape)
(2, 2) (2, 2) (2, 100) (2, 100)
fig, ax = plt.subplots()
fig.set_figheight(8)
fig.set_figwidth(8)
ax.scatter(*Xp.T, alpha=0.3)
ax.scatter(*Xd, color='green', alpha=0.5)
_ = ax.quiver(*Xp.T, *(Xd-Xp.T), alpha=0.2, linestyle='dashed',
linewidth=.4, color='black') # start, end-start
Can be used for compressing and reconstructing the data using U up to k components: $$\underbrace{\mathbf{x}^T}_{\text{rec}} = \underbrace{\underbrace{\mathbf{U}_{|k}}_{k \mapsto D}\quad\underbrace{\underbrace{\mathbf{U}_{|k}^T}_{D \mapsto k}\quad\underbrace{\mathbf{x}^T}_{\text{orig}}}_{projection}}_{reconstruction}$$
Even linear dimensionality reduction is powerful. Here, in uncovers the geography of European countries from only DNA data
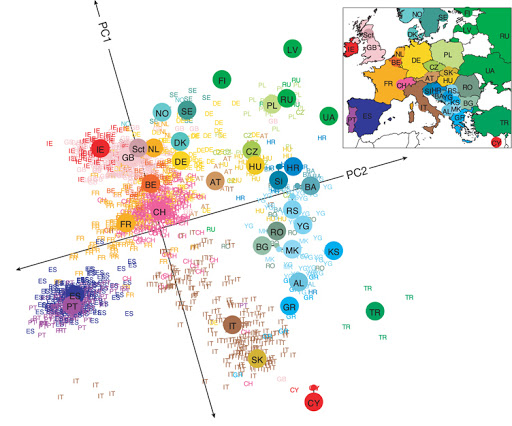
########################################
plt.figure(figsize=(8,8))
plt.scatter(*X.T, alpha=0.3)
plt.scatter(0, 0, c='red')
plt.ylabel('Enjoyment')
plt.xlabel('Skills')
plt.axis('scaled')
plt.xlim(-6, 6)
plt.ylim(-6, 6)
# Taking reconstructed data and shift it back
###############################
Xd_back = (Xd.T*std)+center
#########################
plt.scatter(*Xd_back.T, color='green', marker='.')
<matplotlib.collections.PathCollection at 0x7fd3e18fe760>
conda install pillow or pip install pillowfrom PIL import Image
import requests
from io import BytesIO
#response = requests.get('https://cdn-icons-png.flaticon.com/512/24/24335.png')
img = Image.open('figs/italy.png')
im = np.array(img)
img
X, Y = np.where(im != 0)
sampling = 100 # to have less points
X, Y = X[::sampling], Y[::sampling]
fig = plt.scatter(X, Y, c=X, marker='o', cmap='jet')
plt.scatter(0, 0, c='red')
_ = plt.axis('scaled')
pts = np.stack((X, Y), axis=1)
# Nx2
print(f'num of points {pts.shape[0]} in dimension {pts.shape[1]}' )
num of points 746 in dimension 2
# Standardize the data
# (x-mu)/sigma
center = pts.mean(axis=0)
std = pts.std(axis=0)
pts_z = (pts - center)/std
fig = plt.scatter(*pts_z.T, c=X, marker='.', cmap='jet')
plt.scatter(0,0,c='red')
_ = plt.axis('scaled')
# np.cov wants features on rows
cov = np.cov(pts_z, rowvar=False)
Sigma, U = np.linalg.eig(cov)
# plot principal components aka eigenvectors
fig = plt.scatter(*pts_z.T, c=X, marker='.', cmap='jet')
plt.scatter(0,0,c='red')
plotVectors(fig.axes, [Sigma[0]*U[:,0], Sigma[1]*U[:,1]], cols=['green','blue'])
_ = plt.axis('scaled')
# rotate points
rot = U.T@pts_z.T
np.cov(rot,rowvar=True)
array([[ 3.95280931e-01, -2.38437160e-18],
[-2.38437160e-18, 1.60740363e+00]])
fig, ax = plt.subplots(1,2)
fig.set_figheight(12)
fig.set_figwidth(12)
# First
ax[0].scatter(*pts_z.T, c=X, marker='.', cmap='jet')
ax[0].scatter(0,0,c='red')
plotVectors(ax[0], [Sigma[0]*U[:,0], Sigma[1]*U[:,1]], cols=['green','blue'])
ax[0].set_aspect('equal')
ax[0].set_xlim(-3,3)
ax[0].set_ylim(-3,3)
# second
ax[1].scatter(*rot, c=X, marker='.', cmap='jet')
ax[1].scatter(0,0,c='red')
plotVectors(ax[1], [Sigma[0]*np.array([1,0]), Sigma[1]*np.array([0,1])], cols=['green','blue'])
ax[1].set_aspect('equal')
ax[1].set_xlim(-3,3)
ax[1].set_ylim(-3,3)
(-3.0, 3.0)
np.set_printoptions(suppress=True) #suppress scientific notation, please watch out using this
np.cov(rot, rowvar=True) # rowvar is True because matrix is 2xN
# do you know why we get this? and what are the values inside?
array([[ 0.39528093, -0. ],
[-0. , 1.60740363]])
What does it mean whitening or sphering?
Sigma_inv_sqrt = np.diag(Sigma**-0.5)
# data is rotated and decorrelated (whitening or sphering)
rot_withening = Sigma_inv_sqrt @ U.T @ pts_z.T
fig, ax = plt.subplots(1,3)
fig.set_figheight(14)
fig.set_figwidth(14)
# First
ax[0].scatter(*pts_z.T, c=X, marker='.', cmap='jet')
ax[0].scatter(0,0,c='red')
plotVectors(ax[0], [Sigma[0]*U[:,0], Sigma[1]*U[:,1]], cols=['green','blue'])
ax[0].set_aspect('equal')
ax[0].set_xlim(-3,3)
ax[0].set_ylim(-3,3)
# second
ax[1].scatter(*rot, c=X, marker='.', cmap='jet')
ax[1].scatter(0,0,c='red')
plotVectors(ax[1], [Sigma[0]*np.array([1,0]), Sigma[1]*np.array([0,1])], cols=['green','blue'])
ax[1].set_aspect('equal')
ax[1].set_xlim(-3,3)
ax[1].set_ylim(-3,3)
# third
ax[2].scatter(*rot_withening, c=X, marker='.', cmap='jet')
ax[2].scatter(0,0,c='red')
plotVectors(ax[2], [np.array([1,0]),
np.array([0,1])], cols=['green','blue'])
ax[2].set_aspect('equal')
ax[2].set_xlim(-3,3)
ax[2].set_ylim(-3,3)
(-3.0, 3.0)
fig, ax = plt.subplots(1,3); sizeim=40;
fig.set_figheight(sizeim)
fig.set_figwidth(sizeim)
# First
ax[0].scatter(*pts_z.T, c=X, marker='o', cmap='jet')
ax[0].scatter(0,0,c='red')
plotVectors(ax[0], [Sigma[0]*U[:,0], Sigma[1]*U[:,1]], cols=['green','blue'])
ax[0].set_aspect('equal')
ax[0].set_xlim(-3,3)
ax[0].set_ylim(-3,3)
# second
ax[1].scatter(*rot, c=X, marker='o', cmap='jet')
ax[1].scatter(0,0,c='red')
plotVectors(ax[1], [Sigma[0]*np.array([1,0]), Sigma[1]*np.array([0,1])], cols=['green','blue'])
ax[1].set_aspect('equal')
ax[1].set_xlim(-3,3)
ax[1].set_ylim(-3,3)
# third
ax[2].scatter(*rot_withening, c=X, marker='o', cmap='jet')
ax[2].scatter(0,0,c='red')
plotVectors(ax[2], [np.array([1,0]),
np.array([0,1])], cols=['green','blue'])
ax[2].set_aspect('equal')
ax[2].set_xlim(-3,3)
ax[2].set_ylim(-3,3)
(-3.0, 3.0)
We take the decorrelated data points and compute the covariance matrix:
$$\mathbf{X}_p\mathbf{X}_p^T$$if decorrelated should get us:
$$ \begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix} $$and in more dimensions should give us $\mathbf{I}$ matrix (identity matrix)
np.cov(rot_withening, rowvar=True) # rowvar is True becausse matrix is 2xN
array([[ 1., -0.],
[-0., 1.]])
We can also "prove" it that induced decorrelated and "sphered" points:
$$\underbrace{\mathbf{x}^{T\prime}_p}_{\text{dec.}} = {\mathbf{\Sigma}^{-1/2}\mathbf{U}^T}\quad\underbrace{\mathbf{x}^{T\prime}}_{\text{orig}}$$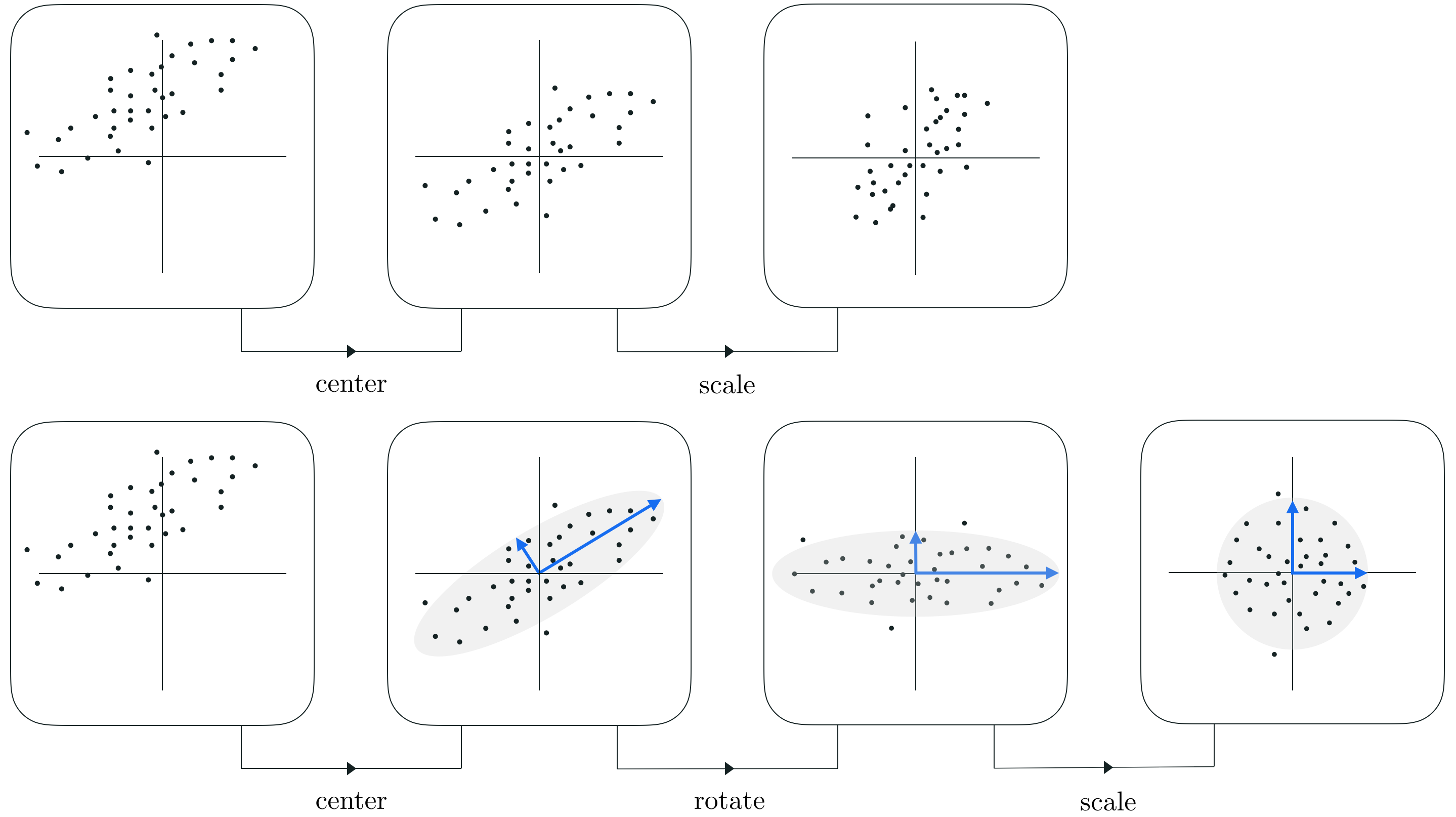
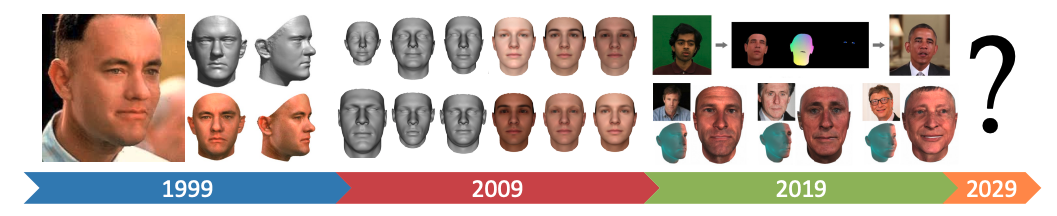
Figure Credit 3DMM, Past Present and Future
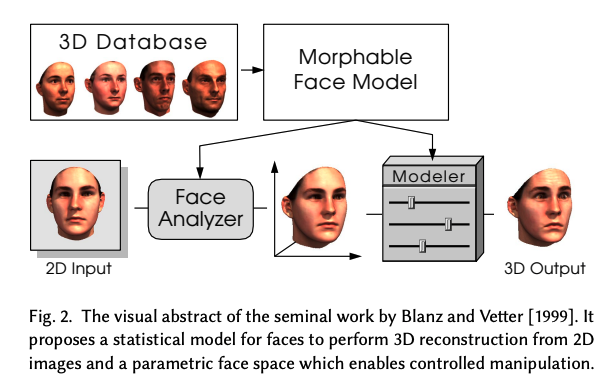
Credits Blanz and Vetter, 1999
$ \mbf{S}^{\prime} = \mbf{\hat{S}} + \sum_{i}^{M-1} \alpha_i \mbf{U}_i $ where:
0.25 * male (1st component) + -0.12 * caucasian (2nd component) + ...$ \mbf{S}^{\prime} = \mbf{\hat{S}} + \sum_{i}^{M-1} \alpha_i \mbf{U}_i $ where:
PCA assumes the data generation process is a multivariate Gaussian distribution (i.e. a Gaussian/Normal distribution but in 3N-D).
$ \underbrace{\{\mathbf{S}_i\}_{i=1}^M}_{\text{known}} \sim \underbrace{\mathcal{N}(\mbf{\hat{S}},\mbf{Cov}_S)}_{\text{multivariate Gaussian assumption}} \approx \underbrace{\mathcal{D}}_{\text{unknown}} $
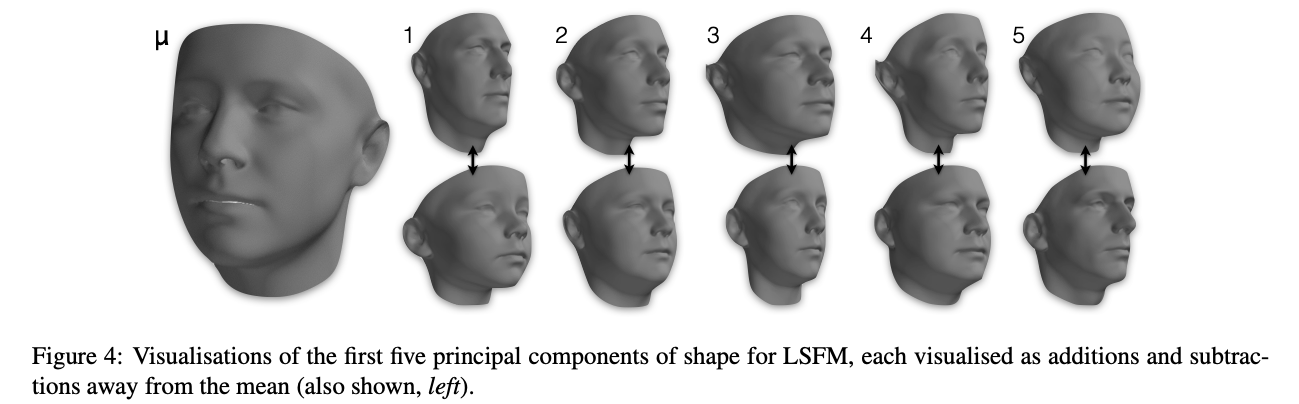
Figure credit Booth et al. 3DMM in the Wild
$\mbf{S}^{\prime} = \mbf{\hat{S}} + \sum_{i}^{M-1} \alpha_i \mbf{U}_i$ where:
Morphable Models are everywhere - Snapfeet App
Body Morphable Models @ Max Planck Institute for Intelligent Systems