

Objective and Motivation: The goal of unsupervised learning is to find hidden patterns in unlabeled data.
\begin{equation} \underbrace{\{\mathbf{x}_i\}_{i=1}^N}_{\text{known}} \sim \underbrace{\mathcal{D}}_{\text{unknown}} \end{equation}Unlike in supervised learning, any data point is not paired with a label.
As you can see the unsupervised learning problem is ill-posed (which hidden patterns?) and in principle more difficult than supervised learning.
Unsupervised learning can be thought of as "finding structure" in the data.
plt.scatter(X[:, 0], X[:, 1], c='black', marker='o', s=50)
plt.show()
plt.scatter(X[:, 0], X[:, 1], c=y, marker='o', s=50, cmap='jet')
plt.show()
from sklearn.datasets import make_blobs
X2, y2 = make_blobs(n_samples=150,
n_features=2,
centers=3,
cluster_std=.7,
shuffle=True,
random_state=0)
plt.scatter(X2[:, 0], X2[:, 1], c='black', marker='o', s=50)
plt.show()
data points 3 maps to the cluster centroid 2plt.scatter(X[:, 0], X[:, 1], c='black', marker='o', s=50)
plt.show()
from sklearn.cluster import KMeans
km = KMeans(n_clusters=3,
init='random',
n_init=10,
max_iter=300,
tol=1e-04,
random_state=0)
y_km = km.fit_predict(X)
plt.scatter(X[y_km == 0, 0],
X[y_km == 0, 1],
s=50,
c='lightgreen',
marker='s',
label='cluster 1')
plt.scatter(X[y_km == 1, 0],
X[y_km == 1, 1],
s=50,
c='orange',
marker='o',
label='cluster 2')
plt.scatter(X[y_km == 2, 0],
X[y_km == 2, 1],
s=50,
c='lightblue',
marker='v',
label='cluster 3')
plt.scatter(km.cluster_centers_[:, 0],
km.cluster_centers_[:, 1],
s=250,
marker='*',
c='red',
label='centroids')
plt.legend()
plt.tight_layout()
plt.show()
Remember that we have to find:
plt.scatter(X[:, 0], X[:, 1], c='black', marker='o', s=50)
plt.show()
Initialization: Random sample the $K$ centroids $\{ \mbf{\mu}_1, \ldots, \mbf{\mu}_k\}$
A trick is using available points (just choose a few of them)
For every point $i=[1,\ldots,N]$ compute the assignment wrt to the $K$ clusters.
#data points x #clusters)Given a point $\mbf{x}_i$, we have to find the label of the "closest" centroids in $\{ \mbf{\mu}_1, \ldots, \mbf{\mu}_k\}$
At the end of this step we should have a pair of aligned set $\{ \mbf{x}_1, \ldots, \mbf{x_n}\}$ and $\{ {y}_1, \ldots, {y_n}\}$
Given a $\mbf{\mu}_k$, we update it as $\mbf{\mu}_k \leftarrow \frac{\sum_i \delta\{y_i=k\}\mbf{x}_i}{\sum_i \delta\{y_i=k\}}$
It is a formalism to express the concept of:
Final result

What does "convergence" mean?
We start from this 150 points in 2-D space
print(X.shape[0])
print(X.shape[1])
150 2
plt.scatter(X[:, 0], X[:, 1], c='black', marker='o', s=50);
plt.show()
def init(X, k):
N, dim = X.shape
# we select k samples from the points we have randomly
idx = np.random.choice(N, k, replace=False)
# the centers are just the k-random sampled points
return X[idx]
def get_assignments(X, centers):
P = X[:, np.newaxis, :] # Nx1xD
diff_sq = (P - centers)**2 # Nx1xD - KxD -> NxKxD # broadcasting
# more info on broadcasting https://numpy.org/doc/stable/user/basics.broadcasting.html
# dist = sum_over_axis(diff^2) then take argmin of dist across centers
y = diff_sq.sum(axis=2).argmin(axis=1)
return y # Nx1 and the values inside go from [1...K]
def update_centers(X, y, K):
# select the points to the k center and remove the N dimension with mean over axis 0
y_row = y.reshape((1, -1)) #1xN
k_col = np.arange(K).reshape(-1, 1) #Kx1
#M[i][j] = 1 if X[j,:] belongs to cluster i else 0
M = k_col == y_row #KxN
#cluster_sum[i,:] contains the sum of all vectors in cluster i
cluster_sum = M @ X #KxN @ NxD -> KxD
#counts[i,:] contains the number of vectors in cluster i
counts = np.sum(M, axis=1).reshape(-1, 1) #Kx1
return cluster_sum / counts #KxD
# Double check
np.random.seed(0) # make the method deterministic
from sklearn.metrics import pairwise_distances_argmin
centers = init(X, 3)
y_assignments = get_assignments(X, centers)
labels = pairwise_distances_argmin(X, centers)
assert np.array_equal(labels, y_assignments), 'Assignment not correct!'
def kmeans(X, n_clusters):
np.random.seed(0) # make the method deterministic
iter = 1
#1. init random sample the centers
centers = init(X, n_clusters)
while True:
# 2. Assignment step
y_assignments = get_assignments(X, centers)
# 3. Update step
new_centers = update_centers(X, y_assignments, n_clusters)
# 4. Check for convergence
# This is a possible implementation
# A more formal is to check that the loss is plateauing (more on this later)
if np.allclose(centers, new_centers):
break
# update cluster for next iter
centers = new_centers
print(f'Iteration {iter}')
iter += 1
return centers, y_assignments
centers, y_est = kmeans(X, 3)
plt.scatter(*X.T, c=y_est,
s=50, cmap='viridis')
_ = plt.scatter(*centers.T,
s=250,
marker='*',
c='red',
label='centroids')
Iteration 1 Iteration 2 Iteration 3
from sklearn.datasets import make_blobs
X, y = make_blobs(n_samples=5000,
n_features=3,
centers=25,
cluster_std=1.25,
shuffle=True,
random_state=0)
fig = plt.figure(figsize=(15,15))
ax = fig.add_subplot(projection='3d')
ax.scatter(*X.T, c=y,
s=50, cmap='jet')
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
_ = ax.set_zlabel('Z Label')
centers, y_est = kmeans(X, 20)
Iteration 1 Iteration 2 Iteration 3 Iteration 4 Iteration 5 Iteration 6 Iteration 7 Iteration 8 Iteration 9 Iteration 10 Iteration 11 Iteration 12 Iteration 13 Iteration 14 Iteration 15 Iteration 16 Iteration 17 Iteration 18 Iteration 19 Iteration 20 Iteration 21 Iteration 22 Iteration 23 Iteration 24 Iteration 25 Iteration 26 Iteration 27 Iteration 28 Iteration 29 Iteration 30 Iteration 31 Iteration 32
fig = plt.figure(figsize=(15,15))
ax = fig.add_subplot(projection='3d')
ax.scatter(*X.T, c=y_est,
s=10, cmap='viridis')
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
_ = ax.set_zlabel('Z Label')
_ = ax.scatter(*centers.T,
s=250,
marker='*',
c='red',
label='centroids')
from sklearn.datasets import make_blobs
from sklearn.metrics import pairwise_distances_argmin
X, y_true = make_blobs(n_samples=300, centers=4,
cluster_std=0.60, random_state=0)
rng = np.random.RandomState(42)
centers = [0, 4] + rng.randn(4, 2)
def draw_points(ax, c, factor=1):
ax.scatter(X[:, 0], X[:, 1], c=c, cmap='viridis',
s=50 * factor, alpha=0.3)
def draw_centers(ax, centers, factor=1, alpha=1.0):
ax.scatter(centers[:, 0], centers[:, 1],
c=np.arange(4), cmap='viridis', s=200 * factor,
alpha=alpha)
ax.scatter(centers[:, 0], centers[:, 1],
c='black', s=50 * factor, alpha=alpha)
def make_ax(fig, gs):
ax = fig.add_subplot(gs)
ax.xaxis.set_major_formatter(plt.NullFormatter())
ax.yaxis.set_major_formatter(plt.NullFormatter())
return ax
fig = plt.figure(figsize=(15, 4))
gs = plt.GridSpec(4, 15, left=0.02, right=0.98, bottom=0.05, top=0.95, wspace=0.2, hspace=0.2)
ax0 = make_ax(fig, gs[:4, :4])
ax0.text(0.98, 0.98, "Random Initialization", transform=ax0.transAxes,
ha='right', va='top', size=16)
draw_points(ax0, 'gray', factor=2)
draw_centers(ax0, centers, factor=2)
for i in range(3):
ax1 = make_ax(fig, gs[:2, 4 + 2 * i:6 + 2 * i])
ax2 = make_ax(fig, gs[2:, 5 + 2 * i:7 + 2 * i])
# E-step
y_pred = pairwise_distances_argmin(X, centers)
draw_points(ax1, y_pred)
draw_centers(ax1, centers)
# M-step
new_centers = np.array([X[y_pred == i].mean(0) for i in range(4)])
draw_points(ax2, y_pred)
draw_centers(ax2, centers, alpha=0.3)
draw_centers(ax2, new_centers)
for i in range(4):
ax2.annotate('', new_centers[i], centers[i],
arrowprops=dict(arrowstyle='->', linewidth=1))
# Finish iteration
centers = new_centers
ax1.text(0.95, 0.95, "E-Step", transform=ax1.transAxes, ha='right', va='top', size=14)
ax2.text(0.95, 0.95, "M-Step", transform=ax2.transAxes, ha='right', va='top', size=14)
# Final E-step
y_pred = pairwise_distances_argmin(X, centers)
axf = make_ax(fig, gs[:4, -4:])
draw_points(axf, y_pred, factor=2)
draw_centers(axf, centers, factor=2)
axf.text(0.98, 0.98, "Final Clustering", transform=axf.transAxes,
ha='right', va='top', size=16)
fig.savefig('figs/05.11-expectation-maximization.png')
/var/folders/rt/lg7n4lt1489270pz_18qn1_c0000gp/T/ipykernel_88826/3638793124.py:11: UserWarning: No data for colormapping provided via 'c'. Parameters 'cmap' will be ignored ax.scatter(X[:, 0], X[:, 1], c=c, cmap='viridis',
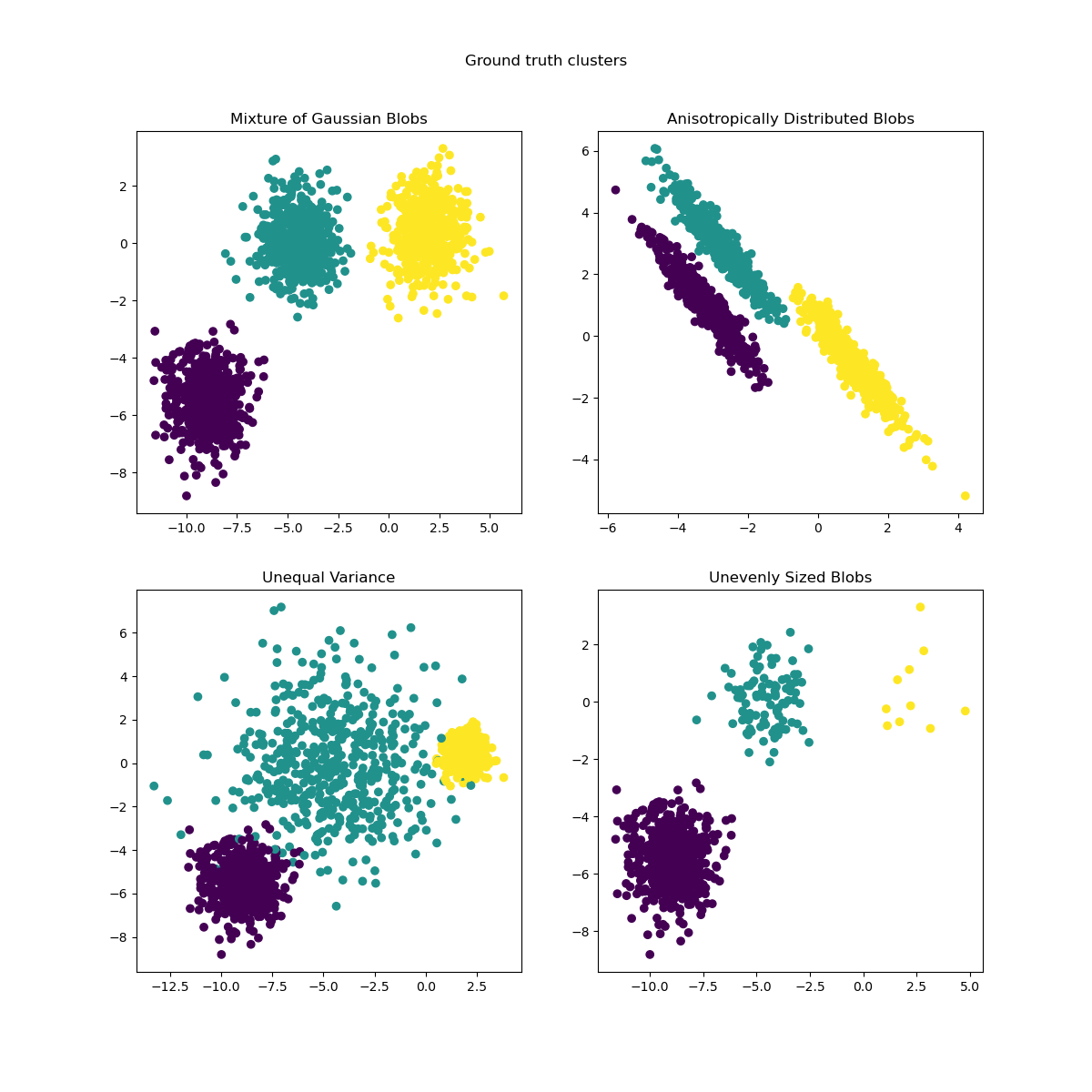
from sklearn.datasets import make_moons
from sklearn.cluster import KMeans
X, y = make_moons(200, noise=.05, random_state=0)
labels = KMeans(2, random_state=0).fit_predict(X)
plt.scatter(X[:, 0], X[:, 1], c=labels,
s=50, cmap='viridis');
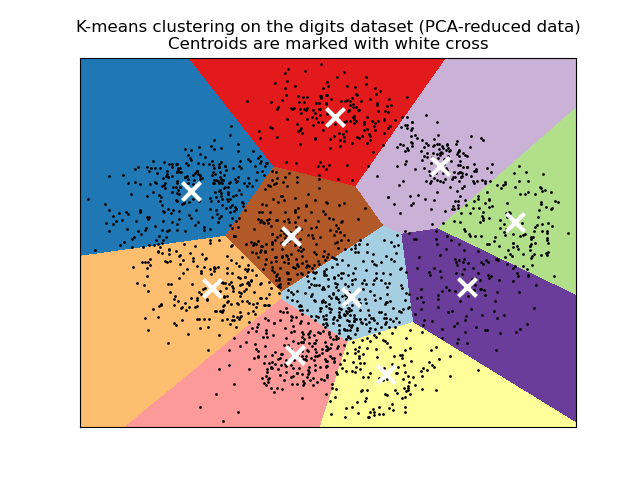
Given a set of points $P_k$ and a distance metric $d(x,y) = \ell_2(x-y)$ and $d(x,\, A) = \inf\{d(x,\, a) \mid a \in A\}$
$$ R_k = \{x \in X \mid d(x, P_k) \leq d(x, P_j)\; \text{for all}\; j \neq k\} $$

Similarly to PCA, we can define a loss or cost function that can better formalize the K-means algorithm.
$$ \mathcal{L}(\mbf{\mu},y;\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \mbf{\mu}_{y_i} \right|\right|_2^2 $$Similarly to PCA, we can define a loss or cost function that can better formalize the K-means algorithm.
$$ \mathcal{L}(\mbf{\mu},y;\mbf{D}) = \sum_{i=1}^N \left| \left| \mbf{x}_i - \mbf{\mu}_{y_i} \right|\right|_2^2 = \sum_{i=1}^N \min_{k \in K} \Big(\left| \left| \mbf{x}_i - \mbf{\mu}_{k} \right|\right|_2^2 \Big) $$Yes, we are guaranteed that K-means will always converge yet to a LOCAL Optimum
For any dataset D and any number of clusters K, the K-means algorithm converges in a finite number of iterations, where convergence is measured by the loss ceasing the change
For any dataset $\mbf{D}$ and any number of clusters $K$, the K-means algorithm converges in a finite number of iterations, where convergence is measured by the loss $\mathcal{L}$ ceasing the change.
There are only two points in which the K-means algorithm changes its values.
We will show that both of these operations can never increase the value of $\mathcal{L}$. Assuming this is true, the rest of the argument is as follows.
It remains to show that the two steps can only decrease the loss.
Looking at a point $i$ suppose that the previous value of $y_i$ is $a$ and the new value is $b$. It must be the case that $|| \mbf{x}_i -\mbf{\mu}_b ||_2^2 \leq || \mbf{x}_i -\mbf{\mu}_a ||_2^2 $. Thus, changing from $a \mapsto b$ can only decrease the loss.
Consider the second form of the loss. The update computes $\mbf{\mu}_k$ as the mean of the data for which $y_i=k$ is precisely the point to minimize squared distance. So, This step too can only decrease the loss.
Though convergence is guaranteed, it does NOT say it is fast (i.e. it may be slow to converge for large datasets).
There are techniques such as mini-batch K-means to speed up convergence.
Practical implications: It will converge but to get a good result you may need either:
A
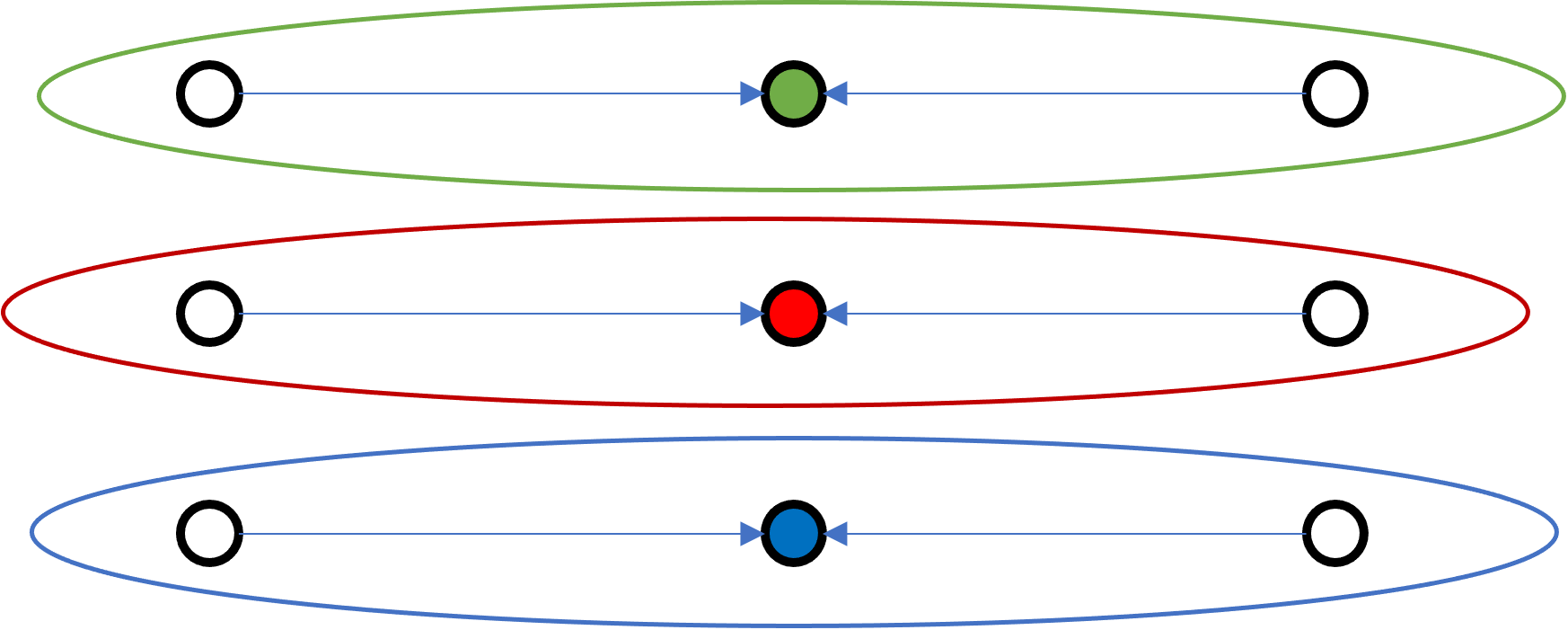
Which one of these configuration is best? A or B?
B
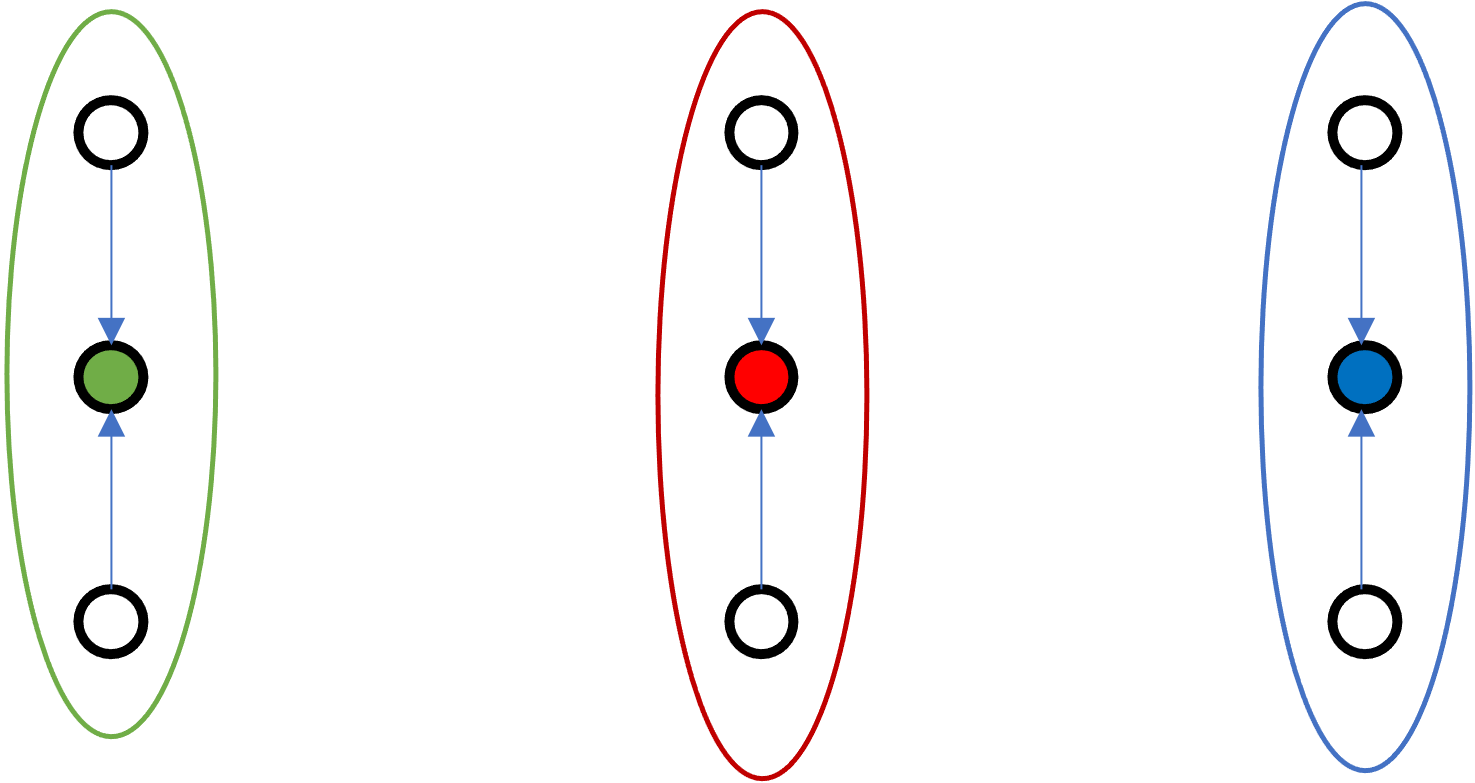
Idea: Semi-random initialization by picking initial means as far from each other as possible
Choose $\mbf{\mu}_1$ arbitrarily between data points
Choose $\mbf{\mu}_1$ arbitrarily between data points
You want the point that’s as far away from all previous means as possible
K-means++ paper idea is pretty "young" (from 2007)
max() across points when selecting the centroidsrandomness(K-means++) > randomness(Kmeans + Furthest-first) because with Furthest-first we select at random only at the start; then all the rest is deterministic. Kmeans++ at each cluster selection, there is stochasticity involveddist = np.array([1.18518298, 1.30917493, 1.10973212, 2.24523519, 1.01625606])
to [0.17262675 0.19068668 0.16163702 0.32702769 0.14802185] simply with
pmf = dist/dist.sum()
fig, axs = plt.subplots(1, 2)
fig.set_figheight(3)
fig.set_figwidth(15)
# PDF
axs[0].stem(pmf, linefmt='b-', markerfmt='bo', basefmt='--')
axs[0].set_title('PMF')
axs[0].set_xlabel('Index of distance')
axs[0].set_ylabel('Probability')
axs[0].set_aspect('auto')
# CUMSUM
axs[1].plot(np.insert(pmf.cumsum(),0,0), 'o--')
axs[1].set_title('CMF')
axs[1].set_xlabel('Index of distance')
axs[1].set_ylabel('Cumulative Probability')
axs[1].set_aspect('auto')
plt.show()
fig, axs = plt.subplots(1, 2)
fig.set_figheight(5)
fig.set_figwidth(20)
# PDF
axs[0].stem(pmf, linefmt='b-', markerfmt='bo', basefmt='--')
axs[0].set_title('PMF')
axs[0].set_xlabel('Index of distance')
axs[0].set_ylabel('Probability')
axs[0].set_aspect('auto')
# CUMSUM
axs[1].plot(np.insert(pmf.cumsum(),0,0), 'o--')
axs[1].set_title('CMF')
axs[1].set_xlabel('Index of distance')
axs[1].set_ylabel('Cumulative Probability')
axs[1].set_aspect('auto')
#axs[1].stem(pmf, linefmt='b-', markerfmt='bo', basefmt='--')
plt.show()
U = np.random.rand(1000) #random sample uniformly from [0, 1].
# u ~ U[0,1]
# for each sampled u.
# find the first bin for which u > cumsum.
# Get the index of that bin as the sample.
sampled_idx = np.argmin((U[:, None] > pmf.cumsum()), axis=1)
import random
random.choices(range(0,5), weights=pmf, k=1000) #support pmf, pmf, k=how many to sample
U = np.random.rand(5)
sampled_idx = np.argmin((U[:, None] > pmf.cumsum()), axis=1)
_=plt.bar(*np.unique(sampled_idx,return_counts=True))
U = np.random.rand(10000)
sampled_idx = np.argmin((U[:, None] > pmf.cumsum()), axis=1)
#_ = plt.hist(sampled_idx, histtype='bar')
_=plt.bar(*np.unique(sampled_idx,return_counts=True))
k that gives a large gap between the k-1-means and the k-means cost function.it is OK for the loss to go down but do not make the model (number of K) too complex.from sklearn.datasets import make_blobs
N_centers = 25
X, y = make_blobs(n_samples=1500,
n_features=2,
centers=N_centers,
cluster_std=.42,
shuffle=True,
random_state=0)
plt.scatter(X[:, 0], X[:, 1], c='black', marker='o', s=50)
plt.show()
def inverse_sampling(pmf, n_samples):
U = np.random.rand(n_samples)
sampled_idx = np.argmin((U[:, None] > pmf.cumsum()), axis=1)
return sampled_idx
def distance(X, centers):
P = X[:, np.newaxis, :] # Nx1xD
diff_sq = (P - centers)**2 # Nx1xD - KxD -> NxKxD # broadcasting
# more info on broadcasting https://numpy.org/doc/stable/user/basics.broadcasting.html
# dist = sum_over_axis(diff^2) then take argmin of dist across centers
diff_sq_sum = diff_sq.sum(axis=2)
return diff_sq_sum
def init(X, K, plusplus=True):
N, dim = X.shape
if not plusplus:
print('> Init with random sampling')
# we select k samples from the points we have randomly
idx = np.random.choice(N, K, replace=False)
# the centers are just the k-random sampled points
return X[idx]
else:
print('> Init with kmeans++')
# we select a point
idx_first = np.random.choice(N, 1)
# init the centers and distances
centers = X[idx_first]
dists = distance(X, centers)
for k in range(1, K): # for [1.....K-1]
# compute min across all k-centers, for all points
# dists is NxK so min_dist is Nx1
min_dist = dists.min(axis=1)
# select one of the N proportional to higher distance
idx = inverse_sampling(min_dist/min_dist.sum(), 1)
new_center = X[idx]
# compute distance between all points to new center
dist_new = distance(X, new_center)
# update pool of centers and distances
centers = np.vstack([centers, new_center])
dists = np.hstack([dists, dist_new])
# the centers are computed after the loop
return centers
def get_assignments(X, centers):
dist_sq = distance(X, centers) # get distance #Nxk
y = dist_sq.argmin(axis=1) # select argmin distance #Nx1
return y, dist_sq.min(axis=1) # Nx1 and the values inside go from [1...K]
def update_centers(X, y, K):
# select the points to the k center and remove the N dimension with mean over axis 0
y_row = y.reshape((1, -1)) #1xN
k_col = np.arange(K).reshape(-1, 1) #Kx1
#M[i][j] = 1 if X[j,:] belongs to cluster i else 0
M = k_col == y_row #KxN
#cluster_sum[i,:] contains the sum of all vectors in cluster i
cluster_sum = M @ X #KxN @ NxD -> KxD
#counts[i,:] contains the number of vectors in cluster i
counts = np.sum(M, axis=1).reshape(-1, 1) #Kx1
return cluster_sum / counts #KxD
# Double check
np.random.seed(0) # make the method deterministic
from sklearn.metrics import pairwise_distances_argmin
centers = init(X, N_centers)
y_assignments, _ = get_assignments(X, centers)
labels = pairwise_distances_argmin(X, centers)
assert np.array_equal(labels, y_assignments), 'Assignment not correct!'
> Init with kmeans++
def kmeans(X, n_clusters):
np.random.seed(0) # make the method deterministic
iter = 1
# 1. init random sample the centers
centers = init(X, n_clusters)
loss = []
while True:
# 2. Assignment step
y_assignments, dist_sq = get_assignments(X, centers)
# distance between points and center, average over points
loss.append(dist_sq.mean())
# 3. Update step
new_centers = update_centers(X, y_assignments, n_clusters)
# 4. Check for convergence
# If two last losses are close enough we stop
if len(loss) > 2 and np.allclose(loss[-2], loss[-1], atol=1e-5):
break
# update cluster for next iter
centers = new_centers
print(f'> Iteration {iter}: loss {loss[-1]}')
iter += 1
return centers, y_assignments, loss
centers, y_est, loss = kmeans(X, N_centers)
> Init with kmeans++ > Iteration 1: loss 1.7296452170971994 > Iteration 2: loss 0.8938248433703356 > Iteration 3: loss 0.7897878328750901 > Iteration 4: loss 0.7802986159291133 > Iteration 5: loss 0.779175352001348 > Iteration 6: loss 0.7782299977540579 > Iteration 7: loss 0.777968597663401 > Iteration 8: loss 0.7778315053744624
fig, ax = plt.subplots(1, 3, figsize=(20, 5))
ax[0].scatter(*X.T, c=y_est,
s=50, cmap='viridis')
ax[0].set_title('Result')
_ = ax[0].scatter(*centers.T,
s=250,
marker='*',
c='red',
label='centroids')
_ = ax[1].plot(loss)
_ = ax[1].set_title('Loss function')
_ = ax[2].scatter(*X.T, c=y,
s=50, cmap='viridis')
_ = ax[2].set_title('Expected')
The applications are inspired from this website
from sklearn.datasets import load_digits
digits = load_digits()
digits.data.shape
(1797, 64)
| Attribute | Value |
|---|---|
| Classes | 10 |
| Samples per class | ~180 |
| Samples total | 1797 |
| Dimensionality | 64 |
| Features | int 0-15 |
_ = plt.hist(digits.data.flatten())
np.random.seed(13) # fixing the seed to see same pictures
fig, ax = plt.subplots(2, 5, figsize=(8, 3))
idx = np.random.choice(digits.data.shape[0], 10)
labels = digits.target[idx, ...]
points = digits.data[idx, ...].reshape(-1, 8, 8)
for axi, vector, y in zip(ax.flat, points, labels):
axi.set(xticks=[], yticks=[])
axi.imshow(vector, interpolation='none', cmap='gray')
axi.set_title(y)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(digits.data,digits.target, test_size=0.1,
random_state=0, stratify=digits.target)
plt.bar(*np.unique(y_train, return_counts=True));
plt.bar(*np.unique(y_test, return_counts=True));
Informally, the mode is the most frequent element
u, counts = np.unique([1,2,3,4,1,1,1,], return_counts=True)
mode = u[counts.argmax()] # will print 1 since 1 is most frequent
# 4 time present
N_centers = 10 # we assume to know that there are 10 digits
centers, y_est, loss = kmeans(X_train, N_centers)
> Init with kmeans++ > Iteration 1: loss 1314.5627705627705 > Iteration 2: loss 756.8259891509574 > Iteration 3: loss 725.4550366992323 > Iteration 4: loss 716.4499767121214 > Iteration 5: loss 711.2049934919781 > Iteration 6: loss 708.236358647983 > Iteration 7: loss 705.2994723513523 > Iteration 8: loss 701.9520972059448 > Iteration 9: loss 698.6658028658622 > Iteration 10: loss 695.5144766870222 > Iteration 11: loss 693.5266489678621 > Iteration 12: loss 692.1208906914004 > Iteration 13: loss 691.3495761746228 > Iteration 14: loss 690.657571369928 > Iteration 15: loss 690.0861205462734 > Iteration 16: loss 689.7669567135049 > Iteration 17: loss 689.6771929699328 > Iteration 18: loss 689.6572694423297 > Iteration 19: loss 689.6474580308259 > Iteration 20: loss 689.6267235993998 > Iteration 21: loss 689.6114085109643
fig, ax = plt.subplots(2, 5, figsize=(8, 3))
centers_img = centers.reshape(10, 8, 8)
for axi, center in zip(ax.flat, centers_img):
axi.set(xticks=[], yticks=[])
axi.imshow(center, interpolation='none', cmap='gray')
KMeans can find clusters whose centers are recognizable digits, with perhaps the exception of 1 and 8.def get_mode(X):
values, counts = np.unique(X, return_counts=True)
return values[counts.argmax()]
# Nx1
labels = np.zeros_like(y_est)
# 10x1
center_label = np.zeros((N_centers), dtype=np.int32)
for i in range(N_centers):
mask = (y_est == i)
mode_y = get_mode(y_train[mask])
labels[mask] = mode_y
center_label[i] = mode_y
from sklearn.metrics import accuracy_score
accuracy_score(y_train, labels)
0.7043908472479901
# sum over axis of dimensionality of squared difference
# Nx1x64 vs 10x64 --> Nx10x64 --sum axis 2 ---> Nx10
# we match the test wrt to the center (there might be other classification methods)
L2_square = ((X_test[...,np.newaxis,:] - centers)**2).sum(axis=2)
# Nx10 -- argmin over axis --> Nx1
idx_cluster = L2_square.argmin(axis=1)
y_test_estimated = center_label[idx_cluster]
accuracy_score(y_test, y_test_estimated)
0.6722222222222223
digits is an easy dataset. Which means distance in the pixel space contains useful information.It is OK to implement things manually in a course but remember that you can use optimized implementation via sklearn.
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=4)
kmeans.fit(X)
y_kmeans = kmeans.predict(X)
t-SNE. Unlike PCA this method supports, multiple non-linear structures so you should see the digits more separated in 2D. For this use sklearn!¶# Note: this requires the ``pillow`` package to be installed
from sklearn.datasets import load_sample_image
china = load_sample_image("china.jpg")
fig, ax = plt.subplots(1, 1, figsize=(20, 15))
ax.set(xticks=[], yticks=[])
ax.imshow(china)
<matplotlib.image.AxesImage at 0x7f9b3152b370>
# Note: this requires the ``pillow`` package to be installed
from sklearn.datasets import load_sample_image
china = load_sample_image("china.jpg")
fig, ax = plt.subplots(1, 1, figsize=(20, 15))
ax.set(xticks=[], yticks=[])
ax.imshow(china)
<matplotlib.image.AxesImage at 0x7f9b40bc81c0>
(427, 640, 3) image WxHx3 where 3 is RGB but can also be seen as a 3D point cloud.data = china/255.0 # scale all points to 0...1
data = data.reshape(-1,3)
data = china/255.0 # scale all points to 0...1
data = data.reshape(-1,3)
data.shape
(273280, 3)
fig = plt.figure(figsize=(15,15))
ax = fig.add_subplot(projection='3d')
data_sampled = data[::10] #1 point every 10 to go faster
ax.scatter(*data_sampled.T, marker='.', color=data_sampled)
ax.set_xlabel('R')
ax.set_ylabel('G')
_ = ax.set_zlabel('B')
ax.view_init(elev=10, azim=(0))
from sklearn.cluster import KMeans
kmeans = KMeans(16)
kmeans.fit(data)
y_est = kmeans.predict(data) # predict cluster center (project)
new_colors = kmeans.cluster_centers_[y_est] # backproject to the colors centers
fig = plt.figure(figsize=(15, 15))
ax = fig.add_subplot(projection='3d')
data_sampled = new_colors[::10] # 1 point every 10 to go faster
ax.scatter(*data_sampled.T, marker='.', color=data_sampled, s=200)
ax.set_xlabel('R')
ax.set_ylabel('G')
_ = ax.set_zlabel('B')
ax.view_init(elev=10, azim=(0))
china_recolored = new_colors.reshape(china.shape)
fig, ax = plt.subplots(1, 2, figsize=(16, 6),
subplot_kw=dict(xticks=[], yticks=[]))
fig.subplots_adjust(wspace=0.05)
ax[0].imshow(china)
ax[0].set_title('Original Image (24 bit per pixel)', size=16) # 8 bit per channel, for each pixel
ax[1].imshow(china_recolored)
ax[1].set_title('16-color Image (4 bit per pixel + 24 bit per centroid)', size=16);