

be the matrix where the columns are the eigenvectors of the matrix $\mathbf{A}$. Let
$$ \boldsymbol{\Sigma} = \begin{bmatrix} 1 & 0 \\ 0 & 4 \end{bmatrix}, $$be the matrix with the associated eigenvalues on the diagonal. Then the definition of eigenvalues and eigenvectors tells us that
$$ \mathbf{A}\mathbf{W} =\mathbf{W} \boldsymbol{\Sigma} . $$If the matrix $\mathbf{W}$ is invertible, we may multiply both sides by $W^{-1}$ on the right, we see that we may write
$$\mathbf{A} = \mathbf{W} \boldsymbol{\Sigma} \mathbf{W}^{-1}.$$One nice thing about eigendecompositions is that we can write many operations we usually encounter cleanly in terms of the eigendecomposition. As a first example, consider:
$$ \mathbf{A}^n = \overbrace{\mathbf{A}\cdots \mathbf{A}}^{\text{$n$ times}} = \overbrace{(\mathbf{W}\boldsymbol{\Sigma} \mathbf{W}^{-1})\cdots(\mathbf{W}\boldsymbol{\Sigma} \mathbf{W}^{-1})}^{\text{$n$ times}} = \mathbf{W}\overbrace{\boldsymbol{\Sigma}\cdots\boldsymbol{\Sigma}}^{\text{$n$ times}}\mathbf{W}^{-1} = \mathbf{W}\boldsymbol{\Sigma}^n \mathbf{W}^{-1}. $$This tells us that for any positive power of a matrix, the eigendecomposition is obtained by just raising the eigenvalues to the same power. The same can be shown for negative powers, so if we want to invert a matrix we need only consider
$$ \mathbf{A}^{-1} = \mathbf{W}\boldsymbol{\Sigma}^{-1} \mathbf{W}^{-1}, $$or in other words, just invert each eigenvalue.
| Method | $\mathbf{A}$ | Decomposition |
|---|---|---|
| SVD | any | $\mathbf{A}=\mathbf{U}\mathbf{\Sigma}\mathbf{V}^T$ |
| Eigen | square | $\mathbf{A}=\mathbf{U}\mathbf{\Sigma}\mathbf{U}^{-1}$ |
| Eigen | square/sym | as above but $\mathbf{U}\mathbf{U}^T=\mathbf{I}$ |
| Method | Step 1 | Step 2 | Step 3 |
|---|---|---|---|
| geometry | rotate | scale axis | rotate |
| SVD | $\mathbf{V}^T$ | $\mathbf{\Sigma}$ | $\mathbf{U}$ |
| geometry | rotate | scale axis | rotate back |
| Eig | $\mathbf{U}^{-1}$ | $\mathbf{\Sigma}$ | $\mathbf{U}$ |
Source: wikipedia

Can be used for compressing and reconstructing the data using U up to k components: $$\underbrace{\mathbf{x}^T}_{\text{prj}} = \underbrace{\underbrace{\mathbf{U}_{|k}}_{k \mapsto D}\quad\underbrace{\underbrace{\mathbf{U}_{|k}^T}_{D \mapsto k}\quad\underbrace{\mathbf{x}^T}_{\text{orig}}}_{projection}}_{reconstruction}$$
$\def\mbf#1{\mathbf{#1}}$
What is the shape of $\mathbb{Cov}$?
We will pull a 2D real dataset made of images in the wild using sklearn
from sklearn.datasets import fetch_lfw_people
faces = fetch_lfw_people(min_faces_per_person=60)
print(faces.target_names)
print(faces.images.shape)
['Ariel Sharon' 'Colin Powell' 'Donald Rumsfeld' 'George W Bush' 'Gerhard Schroeder' 'Hugo Chavez' 'Junichiro Koizumi' 'Tony Blair'] (1348, 62, 47)
faces.images[0,...].shape
(62, 47)
In total we have (1348, 62, 47) so 1348 images of dimension 62x47 = 2914.
# Plot the faces
N_ax, N_img = 10, 10 #10 rows with 10 images per row
fig, ax = plt.subplots(N_ax, N_img,figsize=(10,10),
subplot_kw={'xticks':[], 'yticks':[]},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
for i in range(N_ax):
ax[i,0].set_ylabel(f'Faces {i}')
for j in range(N_img):
ax[i,j].imshow(faces.data[i*N_img+j].reshape(62, 47), cmap='gray')
plt.imshow(faces.data[9*N_img].reshape(62, 47), cmap='gray')
<matplotlib.image.AxesImage at 0x7fd499e50ca0>
# Sampling
samples = np.random.uniform(0, 255, (100, 62, 47)).astype(np.uint8)
# uniformly sample 100 points in 62x47 space.
# Plot the faces
N_ax, N_img = 10, 10 # 10 rows with 10 images per row
fig, ax = plt.subplots(N_ax, N_img, figsize=(10, 10),
subplot_kw={'xticks': [], 'yticks': []},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
for i in range(N_ax):
ax[i, 0].set_ylabel(f'Samples {i}')
for j in range(N_img):
ax[i, j].imshow(samples[i*N_img+j].reshape(62, 47), cmap='gray')
# Sampling
samples = np.random.uniform(0, 255, (100, 62, 47)).astype(np.uint8)
# uniformly sample 100 points in 62x47 space.
# Plot the faces
N_ax, N_img = 10, 10 # 10 rows with 10 images per row
fig, ax = plt.subplots(N_ax, N_img, figsize=(10, 10),
subplot_kw={'xticks': [], 'yticks': []},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
for i in range(N_ax):
ax[i, 0].set_ylabel(f'Samples {i}')
for j in range(N_img):
ax[i, j].imshow(samples[i*N_img+j].reshape(62, 47), cmap='gray')
The phenomenon is also known as...
The number of training samples must grow exponentially with dimensionality if we want to maintain the same "density"
This also means [informally] that if you increase the input dimension, the number of your training samples should grow exponentially to have the same performance and avoid "overfitting".
More on overfitting later but let's say for now that overfitting = ML method not working well
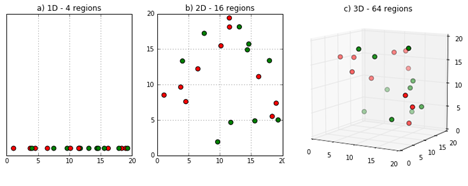
Figure Credit DeepAI
_=plt.imshow(faces.data[0*N_img].reshape(62, 47), cmap='gray')
Faces may be "living" in a lower embedded space defined by:

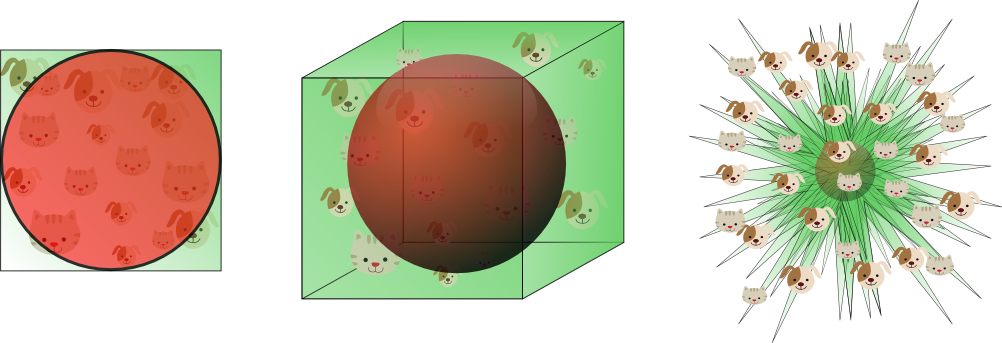
Credit Vincent Spruyt
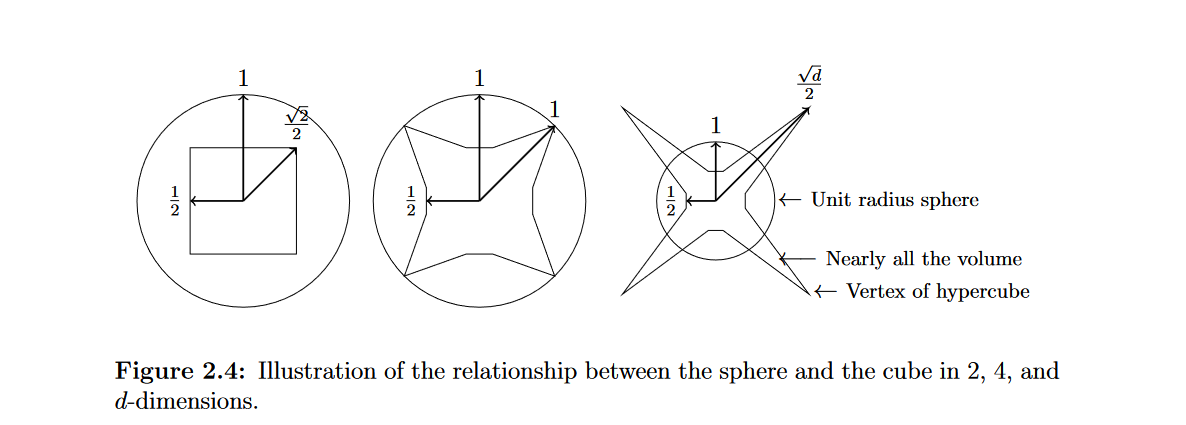
Distances in high dimensions follow a pattern:
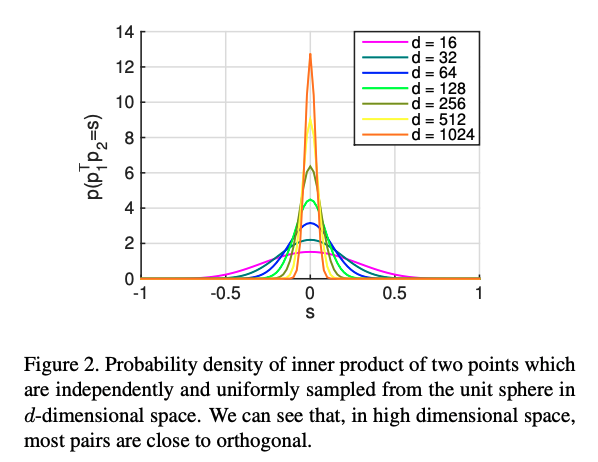
D ≈ 20 is considered “low dimensional,” D ≈ 1,000 is “medium dimensional”D ≈ 100,000 is “high dimensional”.from sklearn.decomposition import PCA
pca = PCA(n_components=150) # retain 150 components
print(faces.data.shape) #NxD N=1348 samples in ~3K-D space
pca.fit(faces.data)
(1348, 2914)
PCA(n_components=150)
fig, axes = plt.subplots(3, 8, figsize=(9, 4),
subplot_kw={'xticks':[], 'yticks':[]},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
for i, ax in enumerate(axes.flat):
if i == 0:
ax.imshow(pca.mean_.reshape(62, 47), cmap='bone') # plot mean
else:
ax.imshow(pca.components_[i].reshape(62, 47), cmap='gray') # plots components
plt.figure(figsize=(20,10))
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
_ = plt.ylim([0,1])
Note: *numpy is not guaranteed to return it ordered so you have to sort*
That is, Keep k components until:
$$\frac{\sum_i^k \lambda_i}{\sum_i^d \lambda_i} \leq 95\%$$plt.rcParams['axes.grid'] = False
from mpl_toolkits.axes_grid1.axes_divider import make_axes_locatable
# Project back to the input
# Project back to the input
projected = pca.transform(faces.data) # project with P = U_t*X_t
unprojected = pca.inverse_transform(projected) # unproject with U*P = U(U_t*X_t)
# Now plot
fig, ax = plt.subplots(3, 10, figsize=(35, 10),
subplot_kw={'xticks': [], 'yticks': []},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
# it is important to get the max value of the error so that
# we plot heatmap error with the SAME SCALE!
errors_img = [(unprojected[i]-faces.data[i])**2 for i in range(10)]
max_val = max([err.max() for err in errors_img])
for i in range(10):
ax[0, i].imshow(faces.data[i].reshape(62, 47), cmap='gray')
ax[1, i].imshow(unprojected[i].reshape(62, 47), cmap='gray')
erri = ax[2, i].imshow((errors_img[i]).reshape(
62, 47), cmap='jet', extent=[0, max_val]*2)
if i == 9:
divider = make_axes_locatable(ax[2, i])
cax = divider.append_axes("right", size="5%", pad=0.005)
plt.colorbar(erri, cax=cax)
ax[0, 0].set_ylabel('full-dim\ninput')
ax[1, 0].set_ylabel('150-dim\nreconstruction')
_ = ax[2, 0].set_ylabel('L2 rec. err')
plt.rcParams['axes.grid'] = False
from mpl_toolkits.axes_grid1.axes_divider import make_axes_locatable
# Project back to the input
# Project back to the input
projected = pca.transform(faces.data) # project with P = U_t*X_t
unprojected = pca.inverse_transform(projected) # unproject with U*P = U(U_t*X_t)
# Now plot
fig, ax = plt.subplots(3, 10, figsize=(35, 10),
subplot_kw={'xticks': [], 'yticks': []},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
# it is important to get the max value of the error so that
# we plot heatmap error with the SAME SCALE!
errors_img = [(unprojected[i]-faces.data[i])**2 for i in range(10)]
max_val = max([err.max() for err in errors_img])
for i in range(10):
ax[0, i].imshow(faces.data[i].reshape(62, 47), cmap='gray')
ax[1, i].imshow(unprojected[i].reshape(62, 47), cmap='gray')
erri = ax[2, i].imshow((errors_img[i]).reshape(
62, 47), cmap='jet', extent=[0, max_val]*2)
if i == 9:
divider = make_axes_locatable(ax[2, i])
cax = divider.append_axes("right", size="5%", pad=0.005)
plt.colorbar(erri, cax=cax)
ax[0, 0].set_ylabel('full-dim\ninput')
ax[1, 0].set_ylabel('150-dim\nreconstruction')
_ = ax[2, 0].set_ylabel('L2 rec. err')
from mpl_toolkits.axes_grid1.axes_divider import make_axes_locatable
############### Fitting with 3 components ######
pca = PCA(n_components=3) # retain 3 components
pca.fit(faces.data)
#############################################
##### Plot
# Project back to the input
projected = pca.transform(faces.data) # project with P = U_t*X_t
unprojected = pca.inverse_transform(projected) # unproject with U*P = U(U_t*X_t)
# Now plot
fig, ax = plt.subplots(3, 10, figsize=(15, 4.5),
subplot_kw={'xticks': [], 'yticks': []},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
# it is important to get the max value of the error so that
# we plot heatmap error with the SAME SCALE!
errors_img = [(unprojected[i]-faces.data[i])**2 for i in range(10)]
max_val = max([err.max() for err in errors_img])
for i in range(10):
ax[0, i].imshow(faces.data[i].reshape(62, 47), cmap='gray')
ax[1, i].imshow(unprojected[i].reshape(62, 47), cmap='gray')
erri = ax[2, i].imshow((errors_img[i]).reshape(
62, 47), cmap='jet', extent=[0, max_val]*2)
if i == 9:
divider = make_axes_locatable(ax[2, i])
cax = divider.append_axes("right", size="5%", pad=0.005)
plt.colorbar(erri, cax=cax)
ax[0, 0].set_ylabel('full-dim\ninput')
ax[1, 0].set_ylabel('3-dim\nreconstruction')
_ = ax[2, 0].set_ylabel('L2 rec. err')
# Nx3
fig = plt.figure(figsize=(5,5))
ax = fig.add_subplot(projection='3d')
_ = ax.scatter(*projected.T, c=faces.target, marker='.', cmap='jet')
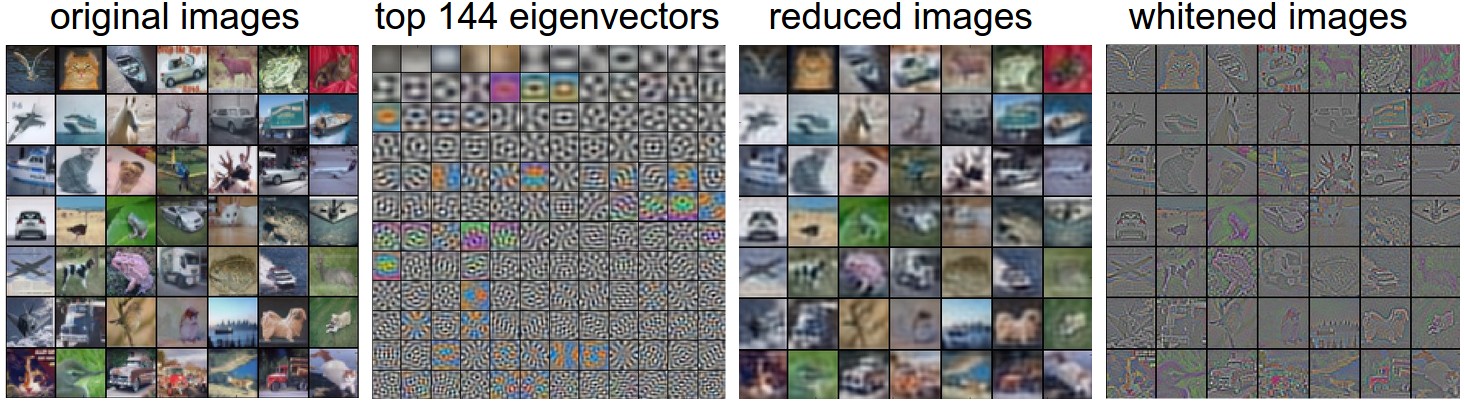

[Taken from cs231n.github.io/neural-networks-2](https://cs231n.github.io/neural-networks-2/)
plt.figure(figsize=(10,10))
np.random.seed(0)
N_samples = 50
# samples points for class 1
X_1 = np.random.uniform(50, 200, N_samples)
X_1 = np.vstack((X_1, (1,)*N_samples))
# samples points for class 2
X_2 = np.random.uniform(50, 200, N_samples)
X_2 = np.vstack((X_2, (20,)*N_samples))
X = np.concatenate((X_1, X_2))
# data
X = np.concatenate((X_1, X_2), axis=1)
# labels
labels = X[1, ...]
# Plot also the training points
plt.scatter(
x=X[0, ...],
y=X[1, ...],
c=labels,
cmap='jet',
)
# Code below wants Nx2
X = X.T
_ = plt.axis('equal')
This example is just used for illustrative purposes since Machine Learning is much related to Information Theory.
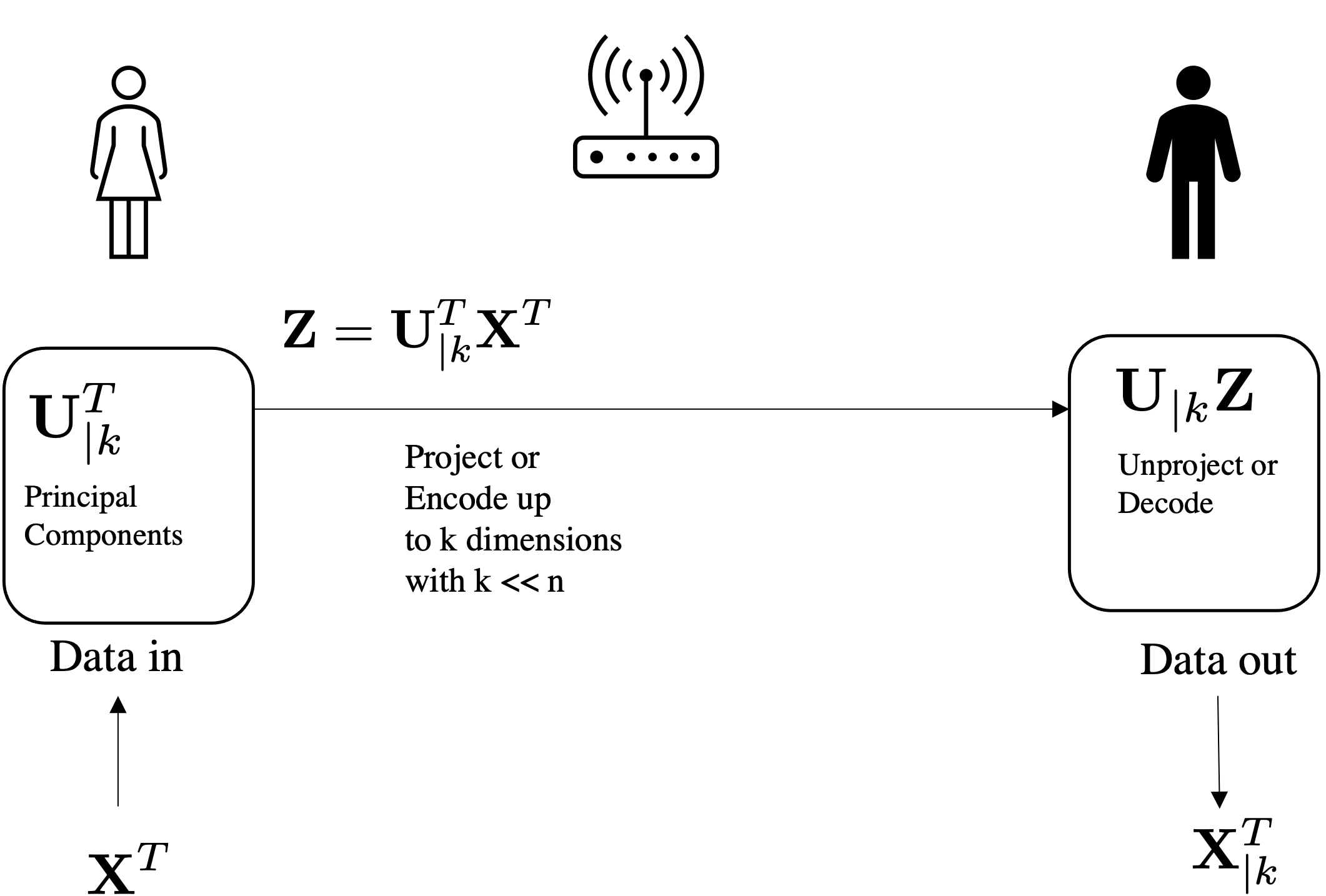

Most of the methods covered in this course are "Inductive" (as opposed to transductive).